A bacterium whose ribosome is made up of proteins containing just 19 amino acids, instead of the 20 used by all life on Earth today, has been engineered by rewriting its genetic code. The work constitutes a key step towards a complete organism that uses fewer amino acids than a natural one – something that could be useful for synthetic biology and provide researchers with the opportunity to study the evolution of organisms with compressed genomes.
Roughly 500 amino acids are found in nature. For reasons that are unclear, 20 of them are used by all organisms in the proteins they use as the core building blocks of their cells. In some rare instances organisms have added a 21st or 22nd amino acid, but no natural organism has functioned with fewer, despite computer modelling suggesting that as few as nine to 12 could hypothetically encode almost all protein folds. Ancient organisms probably used far fewer amino acids than life does today.
The 20 amino acids used to assemble proteins in a cell’s ribosome are encoded by a ‘universal codon table’ comprising 64 possible combinations of three DNA base pairs (codons), many of which code for the same amino acid. Previous work has compressed organisms’ genetic codes by removing some of these redundant triplets. In 2025, researchers in the UK produced Escherichia coli that encoded all 20 amino acids in a 57-codon genetic code, cutting some redundancy in the codon table.
In the new work, the researchers went beyond these ‘synonymous’ mutations and altered E. coli’s genome so that it did not code for the incorporation of one amino acid into the proteins that make up ribosomes. They studied the least conserved amino acids across proteins that performed the same functions in different species, finding that isoleucine was often replaced with another amino acid – most commonly valine – at the same position, so they concluded it might be dispensable. They then introduced genes into E. coli in which the isoleucine in ribosomal proteins was replaced by valine. The organisms survived in some cases, but often they died or didn’t thrive.
The researchers therefore turned to deep learning models of protein structure to work out more sophisticated substitutions. ‘Let’s say nature says “I can replace this isoleucine with a serine, but if I do that I have to replace this other amino acid at this other position because they’re interacting with each other”,’ explains Sergey Ovchinnikov at the Massachusetts Institute of Technology. ‘Protein language models encode all that history inside them.’ Eventually the researchers produced E. coli whose ribosomal proteins contained no isoleucine – although the proteins the ribosomes synthesised for the rest of the cell did. Modified bacterial colonies grew over 60% as quickly as wild type E. coli. ‘We would like to explore not just the ribosome but the entire genome and proteome … to ultimately get to an organism that’s fully stripped down to using only a 19-amino-acid alphabet,’ says Harris Wang at Columbia University, US. ‘I think we’re on a really clear path to achieve this.’
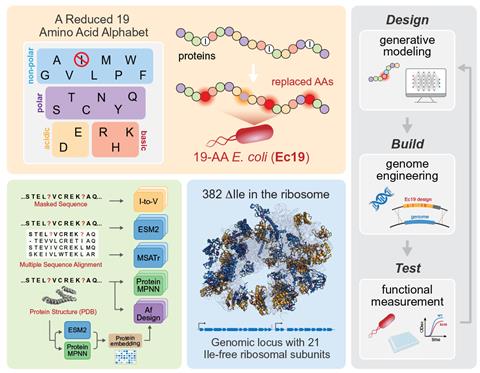
Synthetic biologist Chang Liu of University of California, Irvine says the paper is ‘very significant in giving [the community] a kind of roadmap and a sense of the technical challenges in getting to a 19-amino-acid bacterium’. Organisms without isoleucine – or another amino acid – could have a ‘genetic firewall’ against invasion by viruses that hijack the host’s DNA. ‘If I have a cell that doesn’t use isoleucine, that means I can delete the machinery necessary to incorporate isoleucine into the genetic code.’ This would halt the expression of viral proteins that have isoleucine within them, inactivating the virus. More fundamentally, he says, ‘we could start doing the comparative biology of what a 19-amino-acid protein world would look like … and compare that against our 20-amino-acid world’.
References
L Liu et al, Science, 2026, 392 (DOI: 10.1126/science.aeb5171)














No comments yet