Advances in DNA sequencing technology are changing the way scientists look at genomes. Phillip Broadwith gets up to speed with the latest developments

In April 2003, the Human Genome Project completed a gargantuan task. The international team had toiled for 13 years, at a cost of over £1 billion, to decode the entire sequence of bases in the DNA of a human cell.
Fast forward eight years, and a highly virulent strain of Escherichia coli is spreading through Germany. Accusations fly as to the source of the contamination – is it cucumbers from Spain? Fenugreek seeds from Egypt? Locally produced beansprouts? Pinning down the nature and source of the bacterial strain rapidly and unequivocally becomes crucial for both public health and international political relations.
To do that, researchers turned to the latest in genetic sequencing technology: benchtop machines that can sequence a bacterial genome in a matter of hours.
This is a typical example of how the scientific questions that are being answered using sequencing have changed over the years, explains Julian Parkhill from the Wellcome Trust Sanger Institute in Hinxton, UK – home of the UK end of the Human Genome Project. ‘What we call a genome has changed over time,’ he says. The aim of early sequencing efforts was to produce complete, accurate reference genomes – be they for humans, model organisms or pathogens.
Now, researchers are often more interested in variation between the genomes of individuals. With bacteria, Parkhill says, we can begin looking at how they evolved, how they respond to drugs and vaccines and how they are transmitted. ‘You don’t need [perfect, complete] genomes to answer those questions. You still need them as a reference, but now it’s more about the variation – you need to churn through large numbers of near-identical sequences in order to sift out the very small amounts of variation. In that sense you don’t ever look at a single genome any more.’
Cranking up the pace
To get that amount of information in a realistic timeframe has required huge leaps in the speed and cost of sequencing. ‘We started looking at single bacterial genomes around the time of the Human Genome Project. Around 15 years ago it took maybe two years to do a bacterial genome; in the last two years we’ve sequenced 30,000,’ says Parkhill. ‘Back then it cost something like £500,000; now it’s around £50.’
Cost and throughput – the number of base pairs of sequence an instrument can generate in a given time – have been the main drivers for manufacturers in recent years, fuelled by the goal of the $1000 (£650) human genome. ‘The manufacturers believe, rightly or wrongly, that if you can generate a $1000 human genome, it will open up a whole world of possibilities in personalised medicine,’ says Nick Loman, a bioinformatician from the University of Birmingham, UK.
Loman explains that for the last five years, US firm Illumina has essentially dominated the high-throughput sequencing market. While the underlying chemistry has remained broadly the same, the company has been steadily doubling the throughput of its systems every 6–9 months.
Illumina’s sequencing technology takes a sample of DNA and splits it up into fragments, each around 150 bases long. A short ‘adapter’ sequence of DNA is then added to either end of the fragment, which is complementary to a strand attached to the company’s sequencing flow cell. Using a process similar to the polymerase chain reaction (PCR), each fragment is copied multiple times to make ‘clusters’ of identical fragments attached to the cell. To determine the sequence, the machine introduces fluorescently labelled versions of the four nucleotide bases, and uses a laser and camera to optically determine which clusters have incorporated which fluorescent nucleotides.
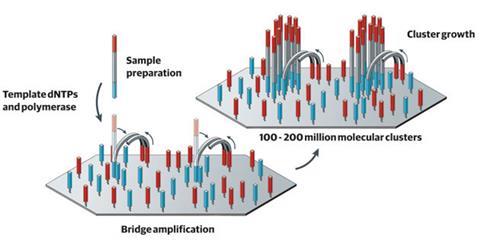
Loman explains that one of the factors that has helped Illumina boost its systems’ performance is that initially the clusters were spaced a long way apart. This meant that when the cell is imaged it would be quite obvious where each cluster started and stopped. ‘There was a worry that if they were too close together, the software wouldn’t be able to work out what was going on, the clusters would grow together, overlap and you’d get a mixed up signal, which would mean you wouldn’t be able to read the sequence,’ Loman adds. ‘But they found that the clusters inhibit each other’s growth, so you can increase the density to get more and more clusters on each chip, but they don’t mix together.’
But cost and throughput are not the only measures of a sequencer’s performance. 454 Life Sciences, now part of Roche’s diagnostics division, has focused on increasing the ‘read length’ of their instruments — the size of the DNA fragments it can sequence. 454 kickstarted high-throughput sequencing with a technology that works similarly to Illumina’s – fragments of DNA are attached to beads and amplified by emulsion PCR. The beads are captured in an array of microscopic wells on a chip, and sequencing is performed by synthesising the complementary strand using fluorescently labelled nucleotides. ‘Read length extension is associated with higher quality,’ says Ben Boese, global marketing manager at 454. ‘Longer reads make it easier to interpret the observations when you do sequencing experiments.’ Whereas Illumina machines currently read fragments of around 150 bases, Boese says the latest 454 machines have read lengths around 700–1000 bases.
Read length becomes particularly important for sequencing organisms that have not been sequenced before, meaning there is no reference genome to work from, says Boese. This is because re-assembling the genome is a bit like putting a jigsaw together – the bigger the pieces the easier the puzzle, especially when some sections of genomes are repeated multiple times, so it can be difficult to be sure where any piece fits. ‘With longer reads, you get a lot more context of the true molecule that was there, and you do a lot less of the statistical inference that might lead you down an incorrect path or leave uncertainty in the results,’ Boese points out.
Cheap as chips
However, the fact that both Illumina and 454’s sequencers rely on optics to capture their data means that storing and processing the data into a sequence requires phenomenal amounts of computer power and storage capacity. This is the bottleneck that US company Ion Torrent is trying to bypass by removing the need for optics entirely.
‘I realised that what we really needed was a purpose-built device that would go from chemical information to digital information directly,’ says Jonathan Rothberg, chief executive of Ion Torrent. The company developed a chip based on a transistor from the 1970s that was sensitive to ions and formed the basis of electronic pH meters. The chip has hundreds of millions of microscopic sensors, Rothberg explains, each capable of sequencing a single strand of DNA as it is replicated. Solutions of each of the four nucleotide bases in turn are repeatedly flooded across the chip. Each time a base is added to a DNA strand, a hydrogen ion is released, which the sensor for that well detects and translates into an electrical signal. This not only removes the need for a camera and a laser, it also means the natural nucleotide bases can be used rather than fluorescently modified ones, which is one of the things that limits the read length of optical-based systems.
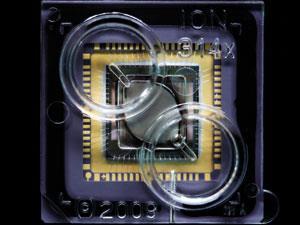

The biggest advantage of using a purely semiconductor-based chip, according to Rothberg, is that it allows the company to tap directly into the trillion-dollar supply chain that serves the electronics industry, rather than having to start from scratch with a totally new technology. ‘If you think about other machines, they’re harder to produce because they were optical instruments that no-one had made before,’ Rothberg says. ‘Our first machines were built in the exact same factories that made Xboxes, so you can use the electronics supply chain for the machine, as well as the semiconductor supply chain for the disposable chips.’
Thanks to this advantage, Ion Torrent’s first generation of chips were ‘a tenth of the cost of the previous generation of technology’, says Rothberg. ‘We think it’s analogous to the introduction of the personal computer, so we called it the personal genome machine.’
As Rothberg points out, semiconductor manufacturing techniques have been advancing in accordance with Moore’s law, which says that the number of transistors that can be fitted on a chip doubles approximately every two years. This means modern day electronics are about 1000-fold more powerful than those manufactured in the 1990s – so by using more modern chip foundries, Ion Torrent can take advantage of the accumulation of Moore’s law and leapfrog directly to more densely packed sensor chips.
The company has just installed its first demonstration models of the Ion Proton sequencer at Baylor College of Medicine in Texas, US. ‘With the Proton, the idea is to make a chip that will do a human genome in about two hours, for $1000,’ says Rothberg. ‘We’re not finished catching up [with the semiconductor industry] yet.’
Technology in the driving seat
All these advances in speed, cost, throughput and read length are not just letting researchers answer the same questions more easily, though. ‘Every time someone brings out a new sequencer that’s got more throughput or longer reads or better error rate, it enables a whole load of experiments that people have been wanting to do for ages,’ says Loman. ‘The technology is completely driving the science.’
A prime example of this is the case of the German E. coli outbreak, which Loman and his colleagues at Birmingham were involved in investigating. Not only did researchers need to identify the strain of bacteria involved, but sequencing it gave them insight into why it was so deadly. ‘At the time there was no reference genome for that strain – there were obviously a lot of other E. colis and most of the genes had been seen before, but never in this particular order,’ Loman says.
That meant they needed the right data to assemble the genome from scratch without making assumptions about the gene order and layout. In doing so, they uncovered several factors contributing to the strain’s lethality, and clues about its evolution and how it might be passed between people. ‘As well as finding Shiga toxin, which is what actually kills you, there are also a number of other virulence factors like aggregative adhesion factors, which may have been responsible for adhering to the gut lumen more tightly and allowing the toxin to transfer into the gut,’ he adds.
Pores to the fore
While most of the latest machines are currently only licensed for research purposes, the prospect of hitting the point where clinical diagnostic sequencing becomes financially viable opens up some exciting possibilities. ‘Imagine you could take blood from a patient with severe sepsis, where every hour counts,’ says Loman. ‘With conventional microbiology you would grow the sample in an aerobic and an anaerobic bottle, you’d see some colonies, then go through a process of trying to identify them, then you’d have to screen for antibiotic resistance. If you could skip all of that and get the identity and the antibiotic resistance profile from the genome in a matter of hours, that would be hugely exciting.’
That kind of application is exactly where UK company Oxford Nanopore believes its soon-to-be-launched sequencers could have the edge. In February, the company announced that it had sequenced its first genome – a small viral genome 5000 base pairs long. While this is a long way from sequencing human genomes, the remarkable aspect was that the company sequenced both strands of the full genome in a single 10 kilobase read, by joining them together into a hairpin structure.
The biggest difference between nanopore sequencing and other high-throughput methods is that it does not rely on synthesising a complementary DNA strand to determine the sequence. Instead, the DNA is fed through a hole in a polymer membrane made by a protein-based nanopore. A second ‘processive’ enzyme controls how fast the DNA ratchets through the pore, and as each base passes through the detector part of the pore, it creates a characteristic difference in the electrical current caused by ions in the surrounding solution flowing through the pore.

John Milton, the company’s chief scientific officer, explained at the time that Oxford Nanopore has spent considerable effort developing the right combination of nanopore and processive enzyme variants, as well as a stable polymer membrane for the pores to sit in, to make the whole system robust. Because it can potentially get a sequence from a single DNA molecule, without needing to make multiple copies to amplify the signal, the company is working towards devices that can handle complex mixed samples like whole blood or very dilute ones like water supplies contaminated with sewage.
But to get the most out of that kind of performance, the sequencer needs to be portable. That’s why, as well as developing a modular lab-based platform, the company unveiled plans for a disposable device called MinION, which will plug into a USB port on a computer.
‘DNA sequencing so far has been fairly specialised,’ says Milton, but he is convinced that new technologies will play a significant role in changing the way people use sequencing. ‘We can provide the shovels, but other people are going to be digging with them,’ he says. ‘If we tried to predict what people will do, I think the only thing we could predict is that our predictions will be wrong.’
Additional information
Animated videos explaining different sequencing methods:
Illumina http://bit.ly/McpzbA
454 Life Sciences http://bit.ly/NwsWcu
Oxford Nanopore http://vimeo.com/36907534
Ion Torrent http://bit.ly/NrdAHt
Comparison of three sequencers in the German E. coli outbreak: N J Loman et al, Nat. Biotechnol., 2012, 30, 434 (DOI: 10.1038/nbt.2198)













No comments yet