New advances that allow scientists to uncover the molecular differences between individual cells could revolutionise medicine, Ian le Guillou finds
As you might expect for someone with a background in chemical physics, Alex Shalek found working with cells during his PhD a frustrating experience. Whether it was killing cancer cells or following the development of immune cells, the results kept varying even though the cells were identical.
‘I was taking what I thought was one system, applying one perturbation, and then getting lots of different outputs,’ says Shalek, now an associate professor of chemistry at Massachusetts Institute of Technology in the US. ‘I was trying to figure out what was happening. One option was that I was just making very bad measurements because I wasn’t trained in biology.’ The more tantalising but unlikely option was that the supposedly identical cells were actually different from one another.
Since 1962, we have known that almost all of the 37 trillion human cells in our bodies contain the same genes. John Gurdon’s Nobel prize-winning experiments with frogs showed that the nucleus from an egg cell could be replaced by one taken from an adult cell, and the egg would grow to become a perfect clone of the original frog.
Whether the DNA comes from a neuron or a liver cell, it contains all the genetic information needed to create every other cell in the body. The difference between these cells isn’t their DNA; it’s how they use it. Activating the right genes at the right time triggers a biochemical cascade that gives each cell the tools it needs to play its role in the body. This pattern of activity varies between cell types and at different stages in their development.
Each time a gene is read, or transcribed, the cell produces a molecule of messenger RNA, which can act as a marker for scientists to track which genes are active. Gathering enough RNA material to analyse, however, has typically required studying bulk samples of thousands of seemingly identical cells. This approach is akin to examining the contents of a fruit smoothie; we can identify all of the molecules that are present, but we have lost the information on which fruit they came from originally and how those fruits differ. What Shalek and his colleagues wanted was a fruit salad, where they could pick out each individual piece and analyse them separately.
It’s a real revolution in resolution
In 2013, they published a study of 18 apparently identical cells of which they had individually sequenced the RNA. ‘When we started doing it, the idea that you could get biologically meaningful data from these sub-picogram levels of RNA in individual cells seemed a little crazy,’ says Shalek. This painstaking work revealed that the cells fell into two distinct subtypes with significant differences in their gene activity despite superficially appearing to be the same. This not only vindicated Shalek’s measuring skills but also demonstrated the need to rethink our categorisation of cells.
A resolution revolution
Together with collaborators from Harvard, MIT and the Broad Institute, Shalek worked to scale up single-cell RNA sequencing. In 2015, this led to Drop-seq, a technique that uses advances in sequencing, microfluidics and molecular barcoding. Each cell is confined to a single droplet in a microfluidic device, where the cells are broken open and the RNA molecules are tagged with a unique identifier for that cell. The droplets can then be pooled and sequenced together, as the molecular barcodes indicate which cell each RNA molecule came from.
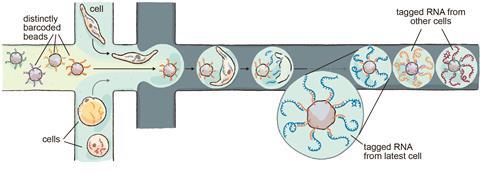
‘It was this incredible coming together of a large number of people around the Boston community. It was one of those things that could only happen in this ecosystem where you had engineers, scientists and computational biologists working together. The concept behind it had been there for a long time, so it was nice to finally see it come to fruition,’ says Shalek.
Just as Robert Hooke used his improved microscope over 350 years ago to see cells for the first time, single-cell RNA sequencing is a tool to see the inner workings of cells and tell them apart – a molecular microscope. ‘It’s a real revolution in resolution,’ says Muzlifah Haniffa, professor of dermatology and immunology at Newcastle University in the UK. Labs around the world have flocked to single-cell RNA sequencing as Drop-seq and other techniques increased the possible number of cells per experiment from hundreds to hundreds of thousands. With many more cells comes the opportunity to identify much rarer cell types that were previously indistinguishable and may be crucially important for understanding health and disease.
Rare cells driving disease
One of the notable successes of single-cell sequencing came in 2018 with the discovery of a new type of lung cell. These pulmonary ionocytes, as they came to be known, have a particularly high activity in the CFTR gene, which is linked to the disease cystic fibrosis. In fact, these rare cells seemed to be responsible for almost all CFTR activity in the lungs, making them a prime target for new treatments.
It’s really about deconstructing the entire disease
Two groups of researchers identified the ionocytes separately and published their findings alongside each other. One of the groups included a collaboration with the Novartis Institute of Biomedical Research (NIBR), where there is still an active programme to characterise these cells.
‘It’s a nice example because it does highlight how powerful it is if you can identify a fairly rare disease-associated cell type,’ says Ulrike Naumann, global lead of next-generation sequencing at NIBR in Basel, Switzerland. ‘It confirmed that NIBR should really be moving into that space and we’ve definitely made some investments since.’
NIBR is using single-cell sequencing to characterise diseases to a new level of detail. For instance, Naumann says, traditional RNA sequencing of many cells struggles to distinguish between different inflammatory skin diseases, as the signal is swamped by an ‘angry keratinocyte [skin cell] signature’. By breaking the diseases down into individual cell types, her team is working to understand why they react differently to treatments.
‘It’s really about deconstructing the entire disease – learn what the critical components are, find those disease-associated cell types, the dysregulated pathways in those cell types and, ultimately, the goal is to find new targets,’ Naumann says.
A periodic table of cells
The ionocytes discovery highlighted the gaps in our knowledge about the importance of rare cell types in how our bodies function. However, filling those gaps is a big undertaking. Haniffa is a member of the Human Cell Atlas initiative, a project that aims to identify all of the different cells in our bodies. Since it was founded in 2016, more than 2000 researchers from 75 countries have joined the initiative.
So far, over a dozen atlases have been published, including for the gut, cerebellum, airways, heart and liver. ‘In the way that you have the periodic table of the elements, what we’ve got is a periodic table of the cells that make up the human body,’ she says.
As if this isn’t ambitious enough, our cells behave very differently over the course of a lifetime, from the earliest stages of development in the womb to old age. Identifying these changes can provide clues into how some diseases develop and find new ways to treat them.
Finding a way to switch off this activity could yield new treatments
Haniffa’s work on a childhood form of kidney cancer called Wilms’ tumour shows the potential value of this approach. Rather than developing from the build-up of mutations over a lifetime like adult cancers, childhood cancers are often caused by a single mutation that disrupts the healthy development of cells. ‘Childhood cancer is often almost like an aberrant form of development,’ she says.
Haniffa found that the tumour had a lot of cells that resemble kidney cells developing in the embryo. It appears that, for some reason, these cancerous cells had stalled in their normal development process. This raises the possibility that treating the tumour cells with growth-signalling molecules could push them to become healthy, replacing the need for chemotherapy to kill the cancer, which can have long-term side effects.
While some diseases, like Wilms’ tumour, can be caused by stalled development, others may be due to developmental processes being triggered in the wrong place and wrong time. Haniffa has found gene activity in diseases such as inflammatory bowel disease or eczema that is normally only found in cells from the developing gut or skin. Finding a way to switch off this programme of activity could yield new treatments for inflammatory diseases.
Understanding infection
The applications to disease go far beyond cancer, though. Even our understanding of infectious diseases could be transformed through single-cell sequencing. The virus that causes Covid-19, Sars-CoV-2, is a notable recent example. Like other viruses, it selectively infects the cells in our body depending on the receptors on their surface.
‘The virus invades the cells and, in an ingenious way, manipulates their molecular machinery,’ says Nikolaus Rajewsky, professor of systems biology at the Max-Delbrück-Center for Molecular Medicine in Berlin, Germany. ‘[Single cell analysis] tells you, for example, which cells of the lung are actually affected, in which way, by the infection or treatment from a molecular point of view.’
The Human Cell Atlas pivoted its focus to Covid-19 when the pandemic hit. It identified particular cells in the eye, nose, salivary glands and lungs that would be susceptible to the Sars-CoV-2 virus. This knowledge helped to inform later research into the different responses between people with Covid-19, as well as provide further evidence on the benefits of masks.
We started applying this technology to everything from Ebola to leprosy
Advances in single-cell technology are helping to expand its reach to infectious disease labs that do not have the infrastructure to carry out techniques like Drop-seq. Shalek, together with Christopher Love, a professor of chemical engineering at MIT, developed a new approach to segregate the cells individually called Seq-Well, which uses what resembles a micro-scale ice cube tray, where each well can hold a single cell.
‘The goal was to remove everything that was complicated,’ says Shalek. ‘We started applying this technology to everything from Ebola to leprosy in various places where there was a real critical need to understand the underlying biology, but barriers to getting these technologies in place, either given infrastructure, costs or accessibility.’
Cell-based medicine
The power of analysing single cells means that we may be able to look beyond treating diseases and instead prevent them from happening altogether. We could detect the earliest changes in cells that deviate from the norm and nudge them back to health, says Rajewsky. ‘The idea of cell-based medicine is to treat cells before symptoms in humans occur.’
Many of the most common age-related health conditions, such as cardiovascular disease, cancer or dementia, take many years or decades to develop. The earliest changes associated with Alzheimer’s disease can start up to 20 or 30 years before the onset of symptoms, by which time there is widespread, irreparable damage to the brain.
Current approaches to these diseases focus on prevention, typically through lifestyle changes, or treatment once the damage is done. Cell-based medicine would fit in the gap between these, treating cells that are starting to go awry before they unravel completely. Single-cell techniques allow these aberrant changes in cells to be detected much earlier than ever before. This could potentially act as a form of health check-up in people who are symptomless, or as a way to monitor those at high risk of specific diseases either through genetic or lifestyle factors. ‘The earliest changes are happening at the single cell level and this is where we can attack,’ says Rajewsky.
The idea of cell-based medicine is to treat cells before symptoms occur
Identifying the first signs of disease can reveal the molecular reasons for these changes and find new targets for treatment. Much like Haniffa’s work on Wilms’ tumour, Rajewsky says that this understanding will allow us to treat damaged cells, rather than simply killing them. One way to correct the cells’ activity would be using engineered mRNA. Just as the Pfizer-BioNTech and Moderna Covid-19 vaccines use mRNA to prompt our cells to produce the Sars-CoV-2 spike protein, mRNA could be used to boost the production of other proteins. ‘mRNA can be programmed to express anything in these cells. So it’s also a wonderful mechanism to interfere with specific cells in a very targeted way,’ says Rajewsky.
Rajewsky is one of the co-ordinators for the LifeTime initiative, a European project across over 90 research institutions using single-cell analyses to advance cell-based medicine. This includes using machine learning techniques to analyse the large amounts of data as well as standardising and streamlining the procedures for studying samples. ‘I think this approach will define the decades to come. It’s not like sequencing the human genome where you have an end product,’ says Rajewsky.
Large-scale, ambitious projects like the Human Cell Atlas or the LifeTime Initiative exemplify the aspirations for single-cell sequencing. However, even if we have a periodic table of cells or can identify the earliest cellular signs of disease, it is still only just the beginning. We are multicellular lifeforms, after all, and every cell is influenced by its neighbours. After breaking down our understanding to the level of individual cells, we will need to learn how to build it back up to dynamic tissues that are in contact with a mixture of cells, both human and microbial.
‘It’s amazing to sit back and think about how far the field’s come in such a short amount of time,’ says Shalek. ‘But as you measure more, it only makes you acutely aware of how much you still have to measure.’
Ian Le Guillou is a science writer based in Paris, France













No comments yet