Philip Ball delves into the suspended animation of cells
Living cells spend a lot of time apparently doing nothing. One of the first things we learn about cells is that they divide and proliferate, but in fact most of the time they don’t. And that’s just how we want it, since – in the human body at least – too much proliferation is dangerous, leading to tumour formation. But as well as the ordinary ‘quiescence’ of healthy cells, some cells are forced into inactivity by lack of the things they need to grow: food, water, warmth. How cells enter and exit these inactive states, and just how inactive they really are, are challenging questions with answers that involve a lot of clever chemistry.
Cells can be regarded as communities of organisms collaborating towards a common end, so shutting down a cell is a little like shutting down a city. If everything just stopped abruptly, the result could be catastrophic. There must be a plan for orderly and gradual shut down, and even in the ‘sleeping’ state some activity has to continue. The cell has to stay ready to reawaken by maintaining some vital stores. And it’s now clear that there is no universal way in which all this happens. Even in our own bodies, different types of cell seem to enter a quiescent, non-dividing phase in different ways. There has been a long debate, still not fully resolved, about whether this quiescence is a passive ‘doing nothing’ or a distinct state that is actively maintained, and whether it is just a slowed down part of the normal cell cycle or a detour from that cycle.
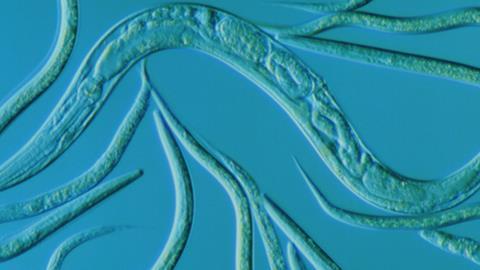
Take, for example, the ‘hibernating’ state of larval parasitic nematode worms such as Caenorhabditiselegans, which can be induced by lack of nutrients or by high temperature. This state is called a dauer (from the German for enduring or persistent), and in this form the worms can increase their lifespan by almost tenfold – as though a human were to go into self-induced suspended animation for many centuries. Dauer larvae are usually motionless, receive no food, and look thin and dense. In other words, they don’t look like ordinary larvae at all, and may not even look alive. In such dormant states, these and other pathogens can be highly resistant to attempts to kill them. Cancer cells can also enter dormancy and become drug resistant, only to reawaken and cause a tumour recurrence.
That’s just one reason why it’s important to understand what goes on when cells shut down. But the more we look, the more questions arise: according to cell biologist Isabelle Sagot of the University of Bordeaux in France, ‘there are a million open questions about cell quiescence’. Researchers aiming to understand these processes are coming to see that the whole notion of being alive is more complex than we might imagine: there are distinct ways of living, of which the usual image of a ‘bag of molecules’ humming with activity is just one.
Heart of glass
In the science fiction version of suspended animation, humans are deep frozen in some cryogenic operation that somehow preserves their tissues intact for reanimation. But the reality may be both simpler and more complicated. As nematode dauers indicate, a dramatic extension of lifespan might not need such complete arrest of all cellular activity, and indeed might not be compatible with it at all. What’s more, ‘freezing up’ of cells might not require cryogenics, but could happen automatically if their metabolism is sufficiently suppressed.
That was the implication of work reported last year by cell biologist Christine Jacobs-Wagner of Yale University in the US and her coworkers.1 They found that the default state of the bacterial cytoplasm – the watery solution of proteins, salts, metabolites and other components that fills bacterial cells – seems to be a kind of glass, in which diffusion of the molecular components is rather slow. In normal conditions, the bacteria avoid entering this sluggish state by actively ‘stirring’ the cytoplasm into a liquid through metabolic processes.
The researchers made this discovery while tracking the movement of a fluorescently labelled protein called crescentin in the bacterium Caulobacter crescentus. Normally crescentin forms filaments on the cell membrane that act as a cytoskeleton. But when given the bulky fluorescent tag, the protein filament detaches and diffuses freely through the cytoplasm. Jacobs-Wagner and colleagues found that if the cells become dormant, with metabolism almost suppressed (as bacteria are apt to do when stressed), the diffusion of crescentin filaments is arrested. Why should such a purely passive physical process be dependent on the metabolic state, they wondered?
Our hypothesis is that cellular activity “shakes” the cytoplasm
Christine Jacobs-Wagner
They then found that small plasmids – rings of DNA – in Escherichia coli also stop diffusing when the cells were depleted of energy. To find similar metabolism-dependent diffusion in bacterial species that diverged over a billion years ago suggests that it is likely to be common in other species. Closer investigation showed that the movements of the molecules is not like Brownian motion in an ordinary liquid. Such differences from normal diffusion have been seen in purely physical systems, such as dense suspensions of colloidal particles close to a glass transition – a switch to a glassy state, in which diffusional motion is more or less arrested even though the material remains non-crystalline. In other words, the cytoplasm was acting rather like a glass.
Jacobs-Wagner and colleagues suspect that, in a low-metabolism state, diffusion in the cytoplasm is hindered by all the other molecular stuff in the way, so that particles get trapped by their neighbours like people in a dense crowd – even though the solvent (water) itself remains fully fluid.
They speculate that metabolic reactions might disrupt this trapping, perhaps in several ways. Metabolic reactions change the charge states or the hydrophobicity (water insolubility) of cytoplasmic molecules, and so can rearrange the shape of the molecular clusters. These reactions often also produce molecular movement that will agitate and stir the cytoplasm and turn it liquid, rather like the way water’s freezing point can be suppressed by constant stirring. ‘Our hypothesis is that cellular activity results in mechanical agitation that “shakes” the cytoplasm,’ says Jacobs-Wagner. Any low metabolism state, whether it is stress-induced dormancy or mere cell quiescence, might then be liable to return to this glassy state, and the largest molecules and particles in the cells would ‘feel’ the sluggishness first. Although this is a purely passive, physical effect rather than an adaptation, it may nevertheless be a beneficial one that ‘preserves’ the cell’s contents in a vitrified matrix.
Crowding of molecules by their neighbours happens in cells even in normal conditions, since the cytoplasm is so densely packed with solutes. In fact, it seems likely that all living cells have evolved not just to cope with crowding but to make active use of it, for example by optimising reaction rates under that constraint so that they’d be less than optimal in dilute solution. However, crowding becomes ever more significant as the cell fluid gets more dense. That’s the case if cells lose water and become dehydrated – a common situation in the most extreme cases of suspended animation, such as spore formation (sporulation) in single-celled organisms. If you think the longevity of dauer worms is impressive, consider that some spores can stay dormant for millions of years. These almost dry cells are pretty much fully vitrified, which makes them highly resistant to all kinds of environmental stresses.
Last year, biophysicist Pascal Hersen of the Paris Diderot University in France and colleagues reported that a reduction in the volume of yeast cells due to osmotic stress – where water is sucked out of the cells by a high external concentration of the sugar alcohol sorbitol – leads to a slowdown of all kinds of biochemical processes, including cell signalling and protein transport in vesicles.2 They figured that this too is caused by extreme crowding, which slows all the dynamics across the board and makes the cytoplasm behave like a colloidal suspension undergoing a glass transition.
Storing up for later
Dormant cells do not, however, seem to be just semi-vitrified versions of active ones. They can look structurally quite different. About 60 years ago, it was discovered that the mitochondria of quiescent yeast cells look very different to those of active cells, forming into dense blobs. Sagot and coworkers have more recently found that yeast cells depleted of a carbon nutrient source strip away their cytoskeleton of actin filaments and store the actin protein in dense clumps that the researchers call actin bodies.3 Under the same conditions, the proteasome, a complex responsible for protein degradation, is moved from the nucleus into cytoplasmic blobs called storage granules. Several other proteins also condense into these clumps as cells enter a quiescent state, and the granules may dissolve when that state is awakened by, for example, adding glucose. In this way, the cells can stockpile their most essential enzymes, saving the energy needed to degrade them while also maintaining reserves so that they can instantly spring back into action when conditions improve.
This isn’t a matter of just sticking the proteins together, says Sagot: the granules tend to be carefully organised. ‘Within actin bodies, actin is very well arranged into filaments,’ she says. Some other quiescent-cell structures, like microtubules bundles (cytoskeletal filaments of tubulin proteins), have ‘very particular architectures’, she adds.
Very recently, Simon Alberti of the Max Planck Institute for Molecular Cell Biology and Genetics in Dresden, Germany, and his coworkers have discovered a particularly striking example of this organised protein storage in dormant cells.4 They looked at yeast under low nutrient conditions, and in particular at what happens to the enzyme glutamine synthetase (Gln1), which catalyses the conversion of glutamate into glutamine – an essential cell reaction. Previous studies have shown that this enzyme, as well as some others, will form filaments in the yeast cells when they are starved of glucose. But why, and how?
Alberti and colleagues found that the trigger for Gln1 filament formation is in fact acidification of the cell, which takes place when nutrient depletion leaves them unable to pump out hydrogen ions. Under acidic conditions, proteins can start to unravel. The formation of filaments in which the enzyme is densely packed might help to shelter and protect them from this process. Gln1 is a decameric protein: it is assembled from 10 identical peptides, which form two rings of five each, stacked face-to-face. Using computer simulations and electron microscopy, Alberti and colleagues found that this stacking can continue back-to-back, so that the decamers line up in chains resembling vertebrae in the spinal column. They think that these chains then pack together to form higher order filament structures – but only in acid conditions, which might change the charge state of the protein surfaces in a way that encourages them to stick together. All this assembly happens only in the crowded environment of the cell. Purified Gln1 in solution formed only the individual stacked chains; these combined into thicker, dense filaments only when the researchers added a polysaccharide to induce crowding.

When clustered into filaments, the enzymes are inactive, perhaps because their shape is changed or because their active sites are obstructed. But Alberti and colleagues found that when they provided glucose to yeast cells that contained Gln1 filaments and which were not able to produce the fresh enzyme themselves from scratch, the cells began to grow and divide again – something possible only if they contained active Gln1, which they could only have got from the disassembled filaments. In other words, the filaments preserve the enzyme for later use, and protect it from the breakdown processes (autophagy) that provide energy in the absence of nutrients. What’s more, cells that were starved under higher pH, so that there was no ‘acid trigger’ for forming Gln1 filaments, subsequently couldn’t grow well – the filaments are clearly needed for surviving the stress.
A pH-triggered entry into a low metabolism state has been seen in several organisms, including hibernating mammals and reptiles. The brine shrimp can survive in dormancy for years this way. Alberti’s group has evidence that not only metabolic enzymes but also many other macromolecular complexes assemble in a pH-dependent manner. ‘We think that a significant fraction of the cytoplasm specifically responds to changes in pH,’ he says. And the assembly is highly selective: only the same type of molecules clump together, via some kind of molecular recognition. Knowing how proteins achieve this to remain stable and viable in the face of stress could be valuable, for example for making protein-based drugs with a long shelf life.
A big turn-off
But even slowing down and making stores isn’t all there is to entering hibernation. Shutting down metabolism is a tricky business. Teymuras Kurzchalia, a specialist in the biochemistry of nematode dauers also at the Max Planck Institute for Molecular Cell Biology and Genetics, compares it to the problem of shutting down a metro network. You can’t just tell all the trains to stop, which would leave them in the wrong places or risk crashes. You need a strategy for ensuring that every vehicle comes to rest safely and in a place that will enable easy start up.
Genetic studies have shown that, as you might expect, quiescent cells express less of the genes associated with cell proliferation. But it’s not enough just to stop dividing. Some genes are actually expressed more during quiescence, and these tend to be ones that prevent the cells from deciding that their arrested division must require some new development – such as undergoing differentiation or death (apoptosis), which may mean shredding their DNA. You might say that cells busily progressing through the cell cycle will interpret the arrest of that cycle as a sign that they need to do something else, unless their gene networks adapt to reassure them that no, it is fine to do nothing but wait for the next signal.
Beyond this, however, it’s hard to generalise about how gene networks respond to quiescence. ‘There is clearly no universal property that has been shown for every quiescent cell type in every quiescence condition,’ says Sagot. ‘I don’t think there’s a single quiescent state that all cells enter when they sense that this is not a time to proliferate,’ agrees cell biologist Hilary Coller of the University of California, Los Angeles in the US. ‘The dormant states of bacteria and yeast have plenty of distinctions from human cellular quiescence.’ Quiescent yeast, for example, accumulates the disaccharide trehalose, both as an energy store and as a protectant against heat, cold and desiccation.
Coller and her coworkers found in 2006 that, even just for human tissue-forming cells called fibroblasts, three different triggers for quiescence resulted in three somewhat different quiescent states.5 Fibroblasts kick into action for healing wounds, but need to become quiescent when they are not required. The researchers arrested the fibroblasts’ proliferation either by depriving them of proteins that trigger division (called mitogens), culturing them at high density, or removing them from the surface on which they adhered. They used an RNA microarray to look at the profiles of gene expression, and found that these differed in each case.
However, there was overlap between the genes that were up- or down-regulated, and nine of them were altered by all of the triggers. So while (for fibroblasts at least) quiescence isn’t a unique state, there does seem to be a central ‘quiescence program’. Some of the genes in this common program have already been identified as tumour suppressors, suggesting that one key to reaching a stable quiescent state is to make sure that interrupting the normal cell cycle isn’t seen as an excuse to evade the usual checks and start proliferating lethally. Another common (although not universal) response is to chill: to reduce metabolic activity and stop producing proteins at the same rate as in the dividing state. Coller thinks that such a program might extend across species, genera and families. ‘There are some attributes of quiescent cells that tend to be conserved from bacteria to yeast to fruitflies to humans,’ she says.
After all, every organism needs some way of shutting its cells down if it is to be resilient in the face of life’s slings and arrows, its gluts and famines. The more we understand about the molecular mechanisms that enable them to hibernate and rest, the better chance we will have of combating the sleeping dangers – the fungal and bacterial spores, the dormant tumour, the parasites biding their time. We won’t necessarily learn how to keep people alive for centuries in suspended animation, but we might be able to deal with some much more immediate problems.
Philip Ball is a science writer based in London, UK
References
1 B R Parry et al, Cell, 2014, 156, 183 (DOI: 10.1016/j.cell.2013.11.028)
2 A Miermont et al, Proc. Natl Acad. Sci. USA, 2013, 110, 5725 (DOI: 10.1073/pnas.1215367110)
3 I Sagot et al, Mol. Biol. Cell., 2006, 17, 4645 (DOI: 10.1091/mbc.E06-04-0282)
4 I Petrovska et al, eLife, in press; or bioRciv DOI: 10.1101/003277
5 H A Coller, L Sang and J M Roberts, PLoS Biol., 2006, 4, e83 (DOI: 10.1371/journal.pbio.0040083); H A Coller, Science, 2011, 334, 1074 (DOI: 10.1126/science.1216242)











No comments yet