New antibiotics could be discovered in future by cracking ‘silent’ genetic codes in bacteria that make unknown natural products. The technique uses computers to predict the chemical structures of suspected antibiotic natural products, which can then be synthesised, offering a much needed route for new drugs to address the ongoing global public health crisis of antibiotic resistance.
It’s estimated that 1.2 million people die of drug-resistant infections every year. Resistance is set to kill 300 million by 2050 and cost the global economy $100 trillion (£80 billion). There is therefore a pressing need to discover drugs that fight pathogens in new ways that bypass existing resistance mechanisms.
Many antibiotics that are in clinical use today originate in bacteria. It’s thought that bacteria evolved the ability to produce antibiotics as part of an arms race against other bacteria in competition for limited resources. As such, their genetics offer a potentially rich seam of potent antibiotic compounds, each encoded by unique clusters of two of more genes.
Bacterial genome sequencing has identified many such biosynthetic gene clusters that encode natural products, and many are thought to be antibiotics. But knowing exactly what these genes make has been a challenge because all too often lab-grown bacteria simply can’t be coerced into producing all of the compounds they are genetically capable of making, resulting in many gene cluster sequences remaining ‘silent’.
In a bid to break this silence and find new antibiotics, Sean Brady’s lab at The Rockerfeller University, US, has now devised a ‘biology-free’ technique that mines libraries of sequenced bacterial genomes to identify and make promising compounds, dubbed synthetic-bioinformatic natural products, or syn-BNPs.
‘Instead of decoding genetic instructions using natural biological processes, bioinformatic algorithms are used to predict the chemical structures produced by bacteria and then chemical synthesis is used to build these potential antibiotics,’ explains Brady.
The team started by searching 10,000 bacterial genomes for a gene that encodes a lipopeptide enzyme, which is commonly seen in gene clusters that produce known lipopeptide antibiotics. Finding 3426 gene clusters with this gene, the researchers then built an evolutionary tree to identify any clusters that were not associated with known lipopeptides. This led them to hone in on a distinct lipopeptide gene cluster in the genome of Paenibacillus mucilaginosus.
Bioinformatic algorithms then predicted eight potential compounds produced by the gene cluster and the team chemically synthesised them. When they tested them for antibacterial activity, the researchers found that one of the new structures, which they called cilagicin, was able to kill multi-drug resistant Gram-positive bacteria.
Modified molecules
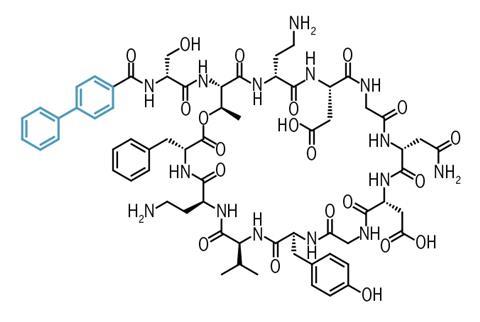
Further experiments revealed a lack of detectable resistance, likely due to cilagicin having two distinct molecular targets that kill bacteria by disrupting cell wall construction, thus making it harder for resistance to evolve. However, when cilagicinwas tested in infected mice, it became apparent that the drug lost its antibacterial clout in the presence of serum, likely because it binds unproductively to serum proteins. The team solved this by modifying the lipid component of cilagicin with biphenyl to produce an analogue, cilagicin BP.
‘Advancements in genomics and in chemistry have allowed scientists to dig deeper than ever before to discover these potent antibiotic weapons, as the Brady group have done with cilagicin,’ says Ian Seiple who investigates chemical synthesis of antibiotics at the University of California, San Francisco. ‘The resulting modified cilagicin BP is a promising starting point for a new class of antibiotics. Generally, multidisciplinary approaches like these are playing a major role in the discovery of next-generation antibiotic candidates,’ adds Seiple.
‘This is an interesting approach that has successfully discovered new antibiotics active against Gram-positive bacteria,’ comments Laura Piddock, scientific director of the Global Antibiotic Research and Development Partnership. ‘I hope that the authors are now applying it to discover new antibiotics active against the Gram-negative bacterial species, pathogens that the World Health Organization has deemed a critical priority for new treatments.’
Earlier work by Brady’s team used the syn-BNP approach to overcome resistance in Gram-negative bacteria to the antibiotic colistin – an antibiotic of last resort – another lipopeptide antibiotic. The technique revealed an improved, naturally occurring analogue of colistin called macolacin, which the researchers further optimised. The approach is not limited to antibiotic discovery either, with Brady’s lab recently reporting the discovery of a potent anti-cancer compound.
‘This is just the tip of the iceberg,’ says Brady. ‘The number of sequenced silent biosynthetic gene clusters already far exceeds the number of characterised natural products. We hope the syn-BNP method can be used to uncover an increasingly diverse collection of new bioactive molecules that are inspired by this large and rapidly growing pool of silent biosynthetic gene clusters.’
References
Z Wang et al, Science, 2022, 376, 991 (DOI: 10.1126/science.abn4213)












No comments yet