Single-cell technique eliminates inefficiencies when identifying immune cells active against specific cancers
A device that screens immune system cells in droplets can select and isolate those that can target specific tumours within days, shaving months off current methods for creating immunotherapies. The system could also be used to personalise cancer treatments.
T cells are white blood cells that attack and eliminate abnormal cells in the body by recognising foreign antigens. T cell receptors, or TCRs, on the T cell surface bind the antigen and activate the T cell against the intruder. Cancerous tumours, however, evade immune detection by releasing immunosuppressive substances and downregulating expression of key proteins of the immune system, among other things. T cell therapy aims to rectify this.
The body contains a huge number of different T cells, but each is specific to a different antigen, which is a problem. In T cell therapy, billions of T cells specific to one tumour antigen enter the body, generating a stronger immune response.
Isolating T cells that express receptors specific to a target antigen is essential for T cell therapy to be successful, but a huge challenge considering the billions of different possible TCRs. ‘The upstream discovery phase is extremely time-consuming and inefficient, and a major roadblock to realising a lot more therapeutic treatments,’ comments Weian Zhao at the University of California Irvine. This is where the device created by Zhao and his colleagues can step in. Normally, it takes months or even years to identify the right therapeutic cells – time many patients do not have – Zhao’s device takes days.
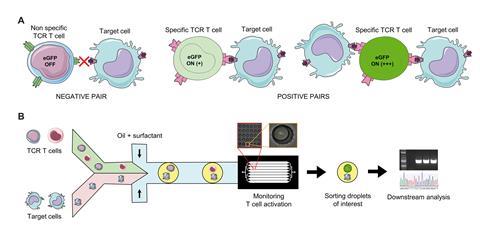
First, the device incubates individual T cells along with target cancer cells in single droplets. The team then monitored the activation kinetics of the T cells by tracking the fluorescence due to expression of eGFP (enhanced green fluorescent protein) in the activated T cells. Other identification techniques rely on binding to identify the desired T cells, meaning they can select false positives – T cells that bind successfully but are not functional. Monitoring activation kinetics in real time allows the scientists to select functional T cells with 100% specificity and be sure they’re not diluted down with non-functional cells. Conventional methods use serial dilutions to achieve a single clone – a time-consuming process. Here, as single cells are analysed, each a single clone, there is no need. Finally, the device analyses a million clones in a single experiment to quickly identify the clones of interest.
‘The reason why this has really attracted a lot of attention and why I think this is so excitingly impactful is because this can really help bring a lot more immunotherapies to the clinic to treat a range of cancers, hopefully at a much lower cost as well,’ comments Zhao. Treatments can currently cost up to half a million dollars or more.
‘This type of technology has the potential to be scaled up and fully standardised for widespread use in the immune-oncology industry,’ comments Rong Fan of Yale University, US, whose research group explores single cell technologies in cancer diagnosis and therapy.
As well as being a tool for immunotherapeutic companies, the device could enable personalised treatment if patient cells are used. ‘This is something we’re really excited about,’ enthuses Zhao, ‘Our goal is to take this to patients.’
Patients and doctors are already paying attention: the team has received enquiries from patients wondering if the technology could help them – can it identify their tumour and the corresponding TCRs they need. For any given patient the technique can profile both their tumour cells as well as their TCRs to identify an ideal patient-specific TCR–antigen match.
In terms of aiding therapeutic discovery, ‘we’re very happy that we’re almost there,’ comments Zhao. ‘As we speak we’re in contact with a number of leading pharmaceutical biotech companies in the world. It’s real, it’s happening.’
References
This article is free to access until 17 May 2019
A Segaliny et al, Lab Chip, 2018, 18, 3733 (DOI: 10.1039/c8lc00818c)








No comments yet