A system based on DNA ‘tiles’ can embody Darwinian evolution, raising new possibilities for understanding natural selection and materials development
‘How Darwin’s “struggle for life” happens has been largely unexplored at the molecular level’, wrote Edith Heard, director-general of the European Molecular Biology Laboratory, recently.1 She was talking about developing a better molecular-scale picture of organisms in their natural environments rather than of bits of them in the lab. But another way to read her comment is that the fundamental processes of Darwinian evolution – replication, mutation, and selection – are not much explored at the level of single molecules.
Of course, in organisms the effects of genetic mutations become visible to selective pressure only at the level of the phenotype. There is a world of complexity separating inherited genotypic mutations from the consequences they have for fitness. But the connection is much more direct in viruses, with consequences of which SARS-CoV-2 is making us painfully aware. And the widespread belief is that Darwinian natural selection began among even simpler molecular entities – perhaps even single molecules, as posited in the RNA world hypothesis for the origin of life.
Templated replication and exponential growth of single DNA-like strands was first demonstrated several decades ago2 and has also been observed in RNA enzymes that can display competition between variants.3 Such a process only becomes Darwinian if there is both mutation and selective pressure that discriminates between mutants. Both of those features have now been demonstrated in synthetic DNA constructs, producing a prototype molecular system that should offer a simple and tunable minimal model of ‘how Darwin’s “struggle for life” happens.’4
His great insight was that DNA can do more than what nature does with it
This new work leverages the DNA nanotechnology pioneered by the late Ned Seeman of New York University, who died in December – and who is a co-author of the paper along with physicist Paul Chaikin and others. Seeman was a one-off: an imaginative visionary, underneath whose gruff and outspoken demeanour was a thoughtful and caring man who inspired great affection from friends and students.5 His great insight was that DNA can do more than what nature does with it – it can serve as a programmable molecule that may self-assemble via base-pairing into all manner of structures and extended arrays. Seeman and others have used synthetic DNA strands to create nanoscale frameworks, lattices, origami patterns and molecular machines.6
Among these possibilities, Seeman, working with Erik Winfree at the California Institute of Technology, made DNA strands that fold into ‘tiles’ that link via their edges into programmable arrays that embody a kind of computation, analogous to the ‘cellular automata’ first posited by Stanislas Ulam and John von Neumann. As it happens, von Neumann conceived of such hypothetical entities as models of self-replication. Seeman, Chaikin and their coworkers have now come full circle, showing that such DNA tiles can indeed embody Darwinian replication and selection – doing in a sense what natural DNA does in genomes, but via very different structures and principles.
The researchers have built on earlier work in which they demonstrated self-replication of the tile patterns.7 The replicating entities here are dimers in which each component is a cross-shaped DNA tile with ‘sticky edges’ where bases can pair up with complementary sequences on other tiles. This pairing is initiated by a seed dimer, which can template the formation of other dimers on the top and bottom edges. The new dimers become covalently united via linking groups activated by UV light, and can themselves template others, producing a growing ladder-like ribbon reminiscent of the double-stranded DNA helix itself. The dimers can then be separated by mild warming, completing the replication cycle.
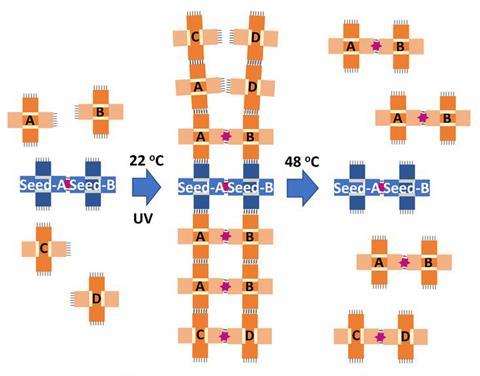
Seeman and colleagues previously showed that this exponential replication can also display selection: with two different ‘species’ of tile, one replicated faster in acidic conditions whereas the other did at neutral pH. Now they have added the other element of Darwinian evolution: mutation. As well as the ‘regular’ (AB) dimer, they introduce another (CD) with sticky-edge sequences similar enough that they can template one another. Then a growing AB ribbon might spawn a CD mutation – and in repeated cycles of replication, the latter may multiply and even take over the population, despite it being seeded initially with an AB dimer. The researchers point out that more mutant types could be added, with different growth rates and sensitivity to environmental factors such as pH or temperature that could act as selective pressures. In this way the system could be used to model experimentally the fundamental features of natural selection – to explore, say, whether there are limits on the range of mutation rates that will support genuine Darwinian evolution, as expected theoretically.
But more than this: the nanoscale building blocks could conceivably be used to create extended materials systems – here in two-dimensional rafts, but others have used DNA constructs to build in three dimensions. That opens up not just replication but natural selection as tools for evolvable materials – an example of the putative field of animate materials that enlist and exploit the capacities of living systems. Seeman’s legacy might be only just beginning.
References
1 E Heard, Nature, 2022, 601, 9 (DOI: 10.1038/d41586-021-03818-3)
2 G von Kiedrowski, Angew. Chem. Int. Ed., 1986, 25, 932 (DOI: 10.1002/anie.198609322)
3 T A Lincoln and G A Joyce, Science, 2009, 323, 1229 (DOI: 10.1126/science.1167856)
4 F Zhou et al, Proc. Natl Acad. Sci USA, 2021, 118, e2111193118 (DOI: 10.1073/pnas.2111193118)
5 P Ball, Nature, 2021, 600, 605 (DOI: 10.1038/d41586-021-03709-7)
6 N C Seeman, Structural DNA Nanotechnology. Cambridge University Press, 2016
7 X He et al., Nat. Mater., 2017, 16, 993 (DOI: 10.1038/nmat4986)











No comments yet