Rachel Brazil talks to the scientists trying to understand the sweet mystery of the glycome
Within our body, simple sugars such as glucose, mannose and fucose connect together to create the complex polysaccharides or glycans that adorn all cell surfaces in a thick coat of sugars. But unlike the genome and the proteome, the glycome is still somewhat a mystery – in part because the chemistry of glycans is so complex. Our bodies synthesise thousands of unique glycans, but there are few analytical methods that can effectively elucidate their structures. And unlike DNA and proteins, which are assembled linearly, glycans are highly branched, making them hard to synthesise. But clever chemistry is providing solutions and bringing us closer to cracking the sugar code.
The oligosaccharides, mostly attached to proteins and lipids, have been the neglected biopolymers of biological research. But Geert-Jan Boons from the University of Georgia in the US says this may be changing. ‘I think over the last 10 years, it has become clear that these carbohydrates are involved in almost every healthy, as well as disease, process.’ Cells’ sugar coatings provide a unique cellular barcode, allowing the immune system to recognise other cells, including foreign invaders. It’s well established that aberrant glycosylation seems to drive cancer progression, specifically hypersialylation – the over-abundance of the nine-carbon backbone sugar sialic acid, found attached to glycan chains’ outermost ends.
The glycome is not directly coded by the genome
The long-term ambition for those in the field is to fully define the glycome – the structure and function of the complete set of glycans produced in a given cell or organism. But this is a little more complicated than defining the genome or proteome, explains Sabine Flitsch, a carbohydrate chemist from the University of Manchester in the UK. ‘The glycome is not directly coded by the genome; you can’t link sugar structure to one particular protein [or gene].’ Glycosylation – the addition of glycans – is a post-translational modification and the over 200 enzymes that assemble sugars do not provide a template, says Boons. ‘[The enzymes] and the availability of sugar nucleotides determines what types of structures are biosynthesised.’
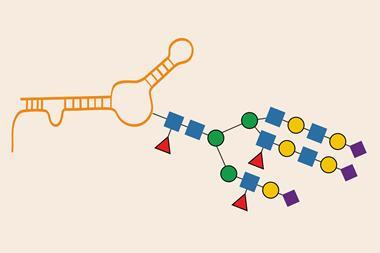
On top of the lack of code, the number of possible glycans is overwhelming. Starting with the 10 common monosaccharides, the potential structures are exponentially greater than those for DNA or proteins. Hexasacchrides, made from six sugar molecules, can adopt 193 billion possible structures. And there are multiple different kinds of glycans , including N -linked oligosaccharides attached to the amino acid asparagine, O -linked oligosaccharides attached to threonine or serine and glycosaminoglycans – long highly polar polysaccharides that act as biological lubricants (an example being heparin, the blood thinner that prevents clots).
Synthetic challenge
Extracting pure glycans from nature is often not possible, so it is chemists who are called upon to synthesise the vast array of possible glycans. But that’s not easy, as they exhibit multiple forms of isomerism. Glycans have numerous arrangement of monosaccharides and variations in branching. Isomers can be based on differing ring sizes, and differing substitution positions (regioisomers). ‘You also [have to] get the stereochemistry of the linkage right – the alpha or beta linkage, which is probably one of the biggest challenges,’ says Flitsch. In glycosylation, a carbohydrate (the glycosyl donor) is attached to a hydroxyl of another sugar (the glycosyl acceptor) creating an anomeric carbon, which allows conversion between two possible configurations. The final product can be in the alpha or beta conformation. ‘You can chemically get selectivity only if you do extensive protection group chemistry,’ explains Flitsch. ‘Each coupling step costs you about seven chemical steps.’
One approach to speeding things up is automation using solid supports. Peter Seeberger, from the Max Planck Institute of Colloids and Interfaces in Potsdam, Germany, founded GlycoUniverse in 2013. The company has created the Glyconeer, an automated oligosaccharide synthesiser, and eight are currently in use around the world.1 ‘The trick in solid state synthesis is that your first building block is attached to a non-soluble polymer, little beads of polystyrene in most cases,’ explains Mario Salwiczek, head of science at GlycoUniverse. ‘And the good thing is that you can wash away all the reactants that have not reacted or all the side products that have formed and you don’t have to do a purification step in between every coupling.’

The longest chains Seeberger has made with the approach are linear 50-sugar oligosaccharide but chains of 6–10 sugars are more typical. ‘Obviously, the technology itself cannot solve the chemistry problems that everybody’s facing,’ Salwiczek notes. One strategy has been to develop generic building blocks of multiple sugars. ‘The solutions that we came up with was clever picking of building blocks,’ says Boons. ‘Oligosaccharides are structurally very complex, but if you look at them carefully, you see certain motifs that are being put together in different ways.’
Boons and others have synthesised such motifs that can then be assembled in multiple ways. Synthetic heparin, made from 20 disaccharides was synthesised this way;2 ‘Those 20 disaccharides were assembled in the end from only six different monosaccharides,’ he says.
Biocatalysis
The other major advance has been the use of glycosyltransferases – the enzymes that transfer monosaccharides from activated sugar mono- or diphosphonucleotides to oligosaccharide chains. In the 1980s the approach was shown to be useful for synthesising complex glycans with each linkage being synthesised by a different enzyme.
What basically took a PhD to synthesise five or six years ago, we do now in the weekend
‘The advantage of these enzymes is that they completely take care of any regio- and stereoselectivity. They take two generic building blocks, and put them together in a very specific way,’ says Flitsch. Both the α- or β-anomers can be made, depending on the nucleotide acceptor. ‘Where you need to have seven steps [with a chemical synthesis], you can do this all in one step,’ adds Flitsch. Chemists are aided in designing syntheses by a database of CAZy enzymes – carbohydrate active enzymes. ‘I don’t say it’s complete,’ explains Flitsch. ‘But we have a fairly good understanding of which enzymes are involved in these biosyntheses.’
‘What basically took a PhD to synthesise five or six years ago, we do now in the weekend,’ says Boons. ‘But the drawback is, if you want to make an unnatural structure and structures that may have more drug-like properties, the enzymes don’t like it.’ He has been trying to get round this by combining chemistry with enzymes. For example, he has synthesised N -glycans with up to four different branches in only 10 chemical and enzymatic steps.
His approach used a symmetric bi-antennary glycosyl derivative building block, isolated from egg yolk powder. He then created an intermediate structure adding an unnatural sugar-nucleotide donor, 5’-disphospho-N -trifluoroacetyl glucosamine. When enzymes are used to further extend the glycan branches, the unnatural molecule is inert to enzymatic action, so further reactions occur selectively at other branches. The unnatural glucosamine is then converted into its natural counterpart, allowing for further enzymatic reactions. Boons calls the strategy ‘stop and go’.3 ‘We really believe that the combination of enzymes and chemistry is revolutionising making these molecules,’ he says.
Analysis in paralysis
Synthesising glycans is only part of the story – it also crucial to be able to analyse them. As well as helping to define the human glycome, it’s important for biopharmaceuticals. Many biological drugs are glycosylated; for example, erythropoietin, the infamous performance-enhancing drug known as EPO, which stimulates red blood cell production. ‘If you don’t glycosylate it, it’s just not active in humans,’ says Flitsch. But trying to analyse the glycan structure is not simple and makes copying biological drugs difficult. ‘The generic [drug] companies are very keen on getting better analytical methods,’ Flitsch adds.
It can be quite challenging to analyse a simple trisaccharide
Most existing analytical techniques have problems when it comes to glycans. Current methods often involve separation techniques interfaced with mass spectrometry. But chromatography protocols were designed with peptides in mind. ‘Sugars are more polar than peptides and as a result, established reverse-phase separation techniques actually do not work properly for sugars,’ says Kevin Pagel, an organic chemist from the Freie Universität Berlin in Germany. The isomerism found with monosaccharides themselves (glucose, galactose and fructose have the same empirical formula) and in their organisation makes mass spectroscopy analysis by mass/charge ratio sometimes impossible. NMR also can’t distinguish stereochemical details. ‘It can be quite challenging to analyse a simple trisaccharide,’ says Pagel.
While N -glycans are better characterised, O -linked glycans are more problematic. Mucins, the large, densely glycosylated proteins with very long glycan chains found in mucus, are a big challenge. Glysoaminoglycans are also hard to analyse, which explains the problems experienced with heparin. In 2008 contaminated heparin caused the deaths of more than 200 people globally due to the manufacturer substituting in a non-active compound, indistinguishable in routine testing.
‘What we really want is something that we can use for everything, that’s sort of universal,’ says Flitsch. ‘At the moment we are at the cusp of being able to do this.’ One technique that may help is ion mobility spectrometry (IMS) – used by airport security to screen for explosives. In combination with mass spectrometry it’s able to distinguish between gas-phase ions based on their mass, charge, size and shape. ‘Different isomers typically have a different size and shape’, Pagel explains, making the method ideal for glycans.
Ions are guided through a gas (helium or nitrogen), under the influence of a weak electric field, and their collision rate will vary, with larger cross-section molecules colliding more frequently and having longer drift times. In tandem with mass spectroscopy, it can distinguish between different isomers of smaller intact glycans.
Fragmenting large glycans before carrying out ion mobility measurements offers something close to a universal sequencing method, using only a limited set of standard fragment fingerprint spectra. ‘It’s sort of like a shotgun sequencing approach,’ says Flitsch. Crucially, carbohydrate fragments seem to retain a memory of their stereochemistry. ‘It will look different, depending if it comes from an alpha or beta linkage – that’s another key step for a sequencing protocol,’ she adds.
There are huge individual differences between people’s glycomes
Another important advance is infrared multiple photon dissociation (IRMPD) spectroscopy. Using a high-intensity laser, an IR spectrum is indirectly measured by observing the disassociation of bonds as they absorb photons. Measuring the fragmentation yield (with a mass spectrometer) as a function of wavelength generates a vibrational spectrum . But there is a drawback: ‘At room temperature the spectral signatures that are obtained are often not very diagnostic because they are typically very broad,’ Pagel explains.
‘What we did for the very first time is to do the experiment with sugars at ultra-cold temperatures,’ says Pagel. ‘We obtained vibrational fingerprints which were incredibly well resolved, which were in fact so well resolved that we were able to disentangle any structural detail in a glycan.’ By cooling the ions in superfluidic helium droplets they reach a temperature of 0.4K and a very low energy state, resulting in a very high resolution spectra.4 Pagel has been able to unambiguously distinguish between a series of trisaccharide isomers that in some cases differed in nothing more than the stereochemistry of a single hydroxyl group.
So combining these new methods, will glycan sequencing become a reality? ‘I think we are beginning to get the synthetic capabilities to pull that off,’ says Boons, although Pagel still thinks sequencing the full human glycome may still be a challenge in 20 years. Unlike DNA and proteins, the glycome is highly dynamic. ‘There are huge individual differences between people’s glycomes,’ notes Flitsch. While DNA differences only occur every thousand residues, the glycome is strongly dependent on environment.
Microarrays
Acknowledging this problem, Lara Mahal from the University of Alberta in Canada, a pioneer in microarray methods for glycan recognition, has a different take on the glycome – she isn’t sure that we actually need it. ‘For a very long time, the community was going after the idea that we needed discreet structures and every single bit of that structure, but if you look at nature, nature doesn’t necessarily care about one discrete structure, nature cares about a collection of structures that do a particular function,’ argues Mahal. ‘The question is, do we need each individual structure [or] can we get a sense of the sub-structural changes?’
To this end Mahal and others are focused on developing microarrays that can study glycan–protein interactions using high-throughput automation. Such arrays arrange various glycans in tiny spots immobilised on a solid support, to be incubated with glycan-binding proteins, cells or even whole viruses. Binding is observed using fluorescent tags.
One of the first methods for immobilising glycans was developed by Ten Feizi at Imperial College London, UK, in the early 2000s. She attached lipid linkers to glycans creating ‘neoglycolipids’, which could then be immobilised to a nitocellulose surface. ‘With covalent array, it’s not very easy to control the density of glycans on the slide,’ says Yan Liu, who heads Imperial’s Carbohydrate Microarray Facility. If glycans are too crowded, or not clustered in the right way, the conditions may not be right for binding. Using neoglycolipids, the glycans are non-covalently bonded to the matrix surface and seem to be able to move and cluster – better mimicking glycan arrangements on cell surfaces. ‘This type of presentation is very good for many types of endogenous and viral glycan binding proteins and very sensitive for antibodies,’ says Liu.
I think it’s probably fair to say carbohydrate chemistry really stretches organic chemistry to its limits
The first method for linking the lipids was reductive amination – a condensation reaction between a primary amine group and the glycan, but this results in ring opening of core monosaccharides, which is a disadvantage for short-chain glycans. In 2007 Liu devised an alternative oxime ligation method.5 The lipid was modified to include an amino-oxy group which was conjugated to form a stable oxime linkage (RHC=NOR′). With this method, ‘we have substantially broadened the scope of the neoglycolipid-based microarray system by providing access to short oligosaccharides and core-branched glycans for ligand-binding studies,’ Liu says.
The glycan library at Imperial is the largest in Europe and has about 1000 sequences – available for all researchers to test against. But there are gaps, with potentially 7000 human recognisable glycans (the actual number isn’t known). ‘I think we’re missing a lot,’ says Liu. ‘On the other hand, we can say we have the major capping groups of these glycans which play important roles for many recognition systems.’
The approach Mahal has taken uses microarrays to find biomarkers for disease, including cancer.6 In the mid-2000s she started to develop high throughput arrays of lectins – carbohydrate-binding plant proteins. ‘Nature has already evolved proteins that have very specific molecular recognition surfaces,’ explains Mahal. ‘They are really good at recognising the difference between, for example, sialic acid that has an alpha-2,6 linkage to a galactose and sialic acid that has an alpha-2,3 linkage. It can tell very immediately those two differences, which is something you cannot tell easily by mass spectrometry.’ Using this approach doesn’t confirm exact structures but ‘it gives us a very rapid way of being able to determine how sugars are changing,’ says Mahal. ‘And that’s really what you care about.’
Mahal’s team have also been trying to understand how glycosylation is controlled in biology and have linked it back to micro-RNA – the small non-coding RNA molecules that silence RNA and regulate post-transcriptional gene expression. Mahal thinks this mechanism controls the transferase enzymes that lead to glycosylation.7 ‘We’re trying to figure out the complete micro-RNA regulation of glycosylation enzymes as a first step towards fixing some of those things,’ says Mahal.
Clearly, both in terms of mapping the glycome and understanding the underpinning biology, there is a long way to go. ‘I think it’s probably fair to say carbohydrate chemistry really stretches organic chemistry to its limits,’ muses Flitsch. But Mahal cautions against seeing glycomics as an insurmountably large scientific mountain. ‘I think one of the problems with glycomics has always been that people have this perception that it is somehow hard, and I would challenge that… because realistically [in biology] what isn’t complex?’
Rachel Brazil is a science writer based in London, UK
References
1 H S Hahm et al,Proc. Natl Acad. Sci. USA, 2017, 114, E3385 (DOI: 10.1073/pnas.1700141114)
2 Q Liang et al,Carbohydr. Res., 2018, 465, 16 (DOI: 10.1016/j.carres.2018.06.002)
3 L Liu et al,Nat. Chem., 2019, 11, 161 (DOI: 10.1038/s41557-018-0188-3)
4 E Mucha et al,Angew. Chem. Int. Ed., 2017, 56, 11248 (DOI: 10.1002/anie.201702896)
5 Y Liu et al,Chem. Biol., 2007, 14, 847 (DOI: 10.1016/j.chembiol.2007.06.009)
6 E L Bird-Lieberman et al,Nat. Med., 2012, 18, 315 (DOI: 10.1038/nm.2616)
7 T Kurcon et al,Proc. Natl Acad. Sci. USA, 2015, 112, 7327 (DOI: 10.1073/pnas.1502076112)










No comments yet