Why did nature stop at 20 amino acids? Rachel Brazil tries to answer the question – and see what it means for how life began and evolved
For many researchers, studying the chemical origins of life is a side project – it’s what they do in between their grant-funded work on the causes and cure of human disease. But understanding evolution at the chemical level is their passion, even when funding is sparse. How chemistry could have brought us to complex life poses many open questions. One fundamental question is why life is based on a set 20 amino acids. Why 20 and not 10 or 30? And why those particular 20? Over the last few decades, the passionate chemists and molecular biologists who can’t leave these questions alone have started piecing together some convincing explanations.
From alanine (A) to tyrosine (Y), 20 ‘proteinogenic’ amino acids, each abbreviated to a different initial, make up the alphabet soup of life. They are the building blocks for proteins, biology’s workhorse macromolecules that provide structure and function in all organisms. But why amino acids? Bernd Moosmann, an expert in redox medicine at the Johannes Gutenberg University of Mainz in Germany suggests the first amino acids were used to anchor membranes to RNA structures: ‘You can see this even in modern life: DNA and RNA in bacteria and mitochondria are always attached from the inside to a membrane.’ Most researcher think this would have been occurring at least 4 billion years ago in an ‘RNA world’, where RNA molecules were the first self-replicators, as well as performing the catalytic role that proteins play today.
How the proteinogenic amino acids came to be on earth is another crucial question. The famous Miller–Urey experiment from 1952 showed that with electric sparks simulating lightning, simple compounds like water, methane, ammonia and hydrogen would form well over 20 different amino acids.1 They are also found in meteorites: analysis of the Murichison meteorite, which landed in Australia in 1969, found at least 86 amino acids, substituent chains of up to nine carbon atoms and dicarboxyl and diamino functional groups.2 Perhaps these generally simple and readily available amino acids were the first to be press-ganged into life?
Andrew Doig, a chemical biologist at the University of Manchester in the UK, has been thinking about the chemistry of evolution, when not carrying out his research into Alzheimer’s disease. He has a different take on the question: ‘[The proteinogenic amino acids] were chosen in the RNA world, where there had been life and metabolism for millions of years, already generating a vast number of organic molecules.’ If amino acids were a product of RNA metabolism this would hugely increase their concentrations in the environment, he argues.
But the selection of the 20 amino acids used in biology is clearly linked to the development of proteins. By polymerising amino acids in long polypeptide chains, proteins could fold into soluble structures with close-packed cores and ordered binding pockets. The arrival of proteins and the eventual adoption of the standard 20 amino acids was likely to have been a big evolutionary step.

But according to Doig, this is all speculation. ‘We have no direct evidence at all.’ What we do know from comparing genomes of organisms today is that by 3.5–3.8 billion years ago our common ancestor – known as the last universal common ancestor – was using the 20 amino acids common to all living things.
A frozen accident?
So why that particular set of 20 amino acids rather than any other? ‘The obvious missing thing is the ability to do redox reactions,’ explains Doig. ‘They weren’t selected for ability to do catalysis directly.’ Today, proteins form enzymes for biological catalysis, but the first biological catalysts in the RNA world were probably what we now call co-factors – metal ions or non-protein organic molecules (coenzymes) that assist enzymes during the catalysis of reactions and are often made from vitamins.
There has been a tendency to see the choice of the 20 amino acids as arbitrary – as in the ‘frozen accident theory’ proposed by British molecular biologist Francis Crick in the 1960s, which suggested a different group of 20 would be just as good. ‘I kept on reading this and realising this wasn’t right,’ says Doig. This spurred him on to put down his thoughts in a recent paper where he argues there are reasons for the selection of every amino acid making them a near ideal group.3 The factors he took into account included each amino acid’s component atoms, functional groups and biosynthetic cost.
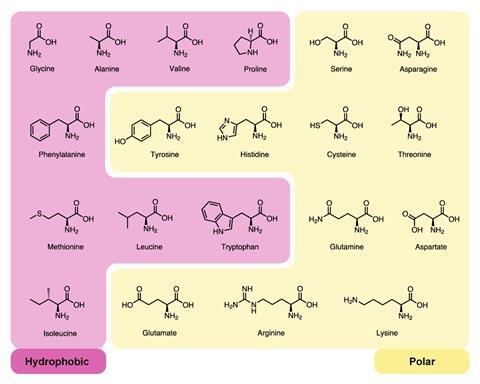
Forming soluble, stable protein structures with close‐packed cores and ordered binding pockets needed the variety of amino acids we see today, explains Doig. Multiple hydrophobic proteins are needed. ‘The core of a protein is a 3D jigsaw puzzle – if you have lots of different hydrophobic amino acids it gives you more options to build a core without any gaps.’
The fact that the hydrophobic amino acids tend to have branched side chains can also be explained. Inside the protein core, the molecule is no longer able to rotate and loses some of its associated entropy. ‘If you have branched amino acids like valine, leucine and isoleucine, you lose less entropy when you bury them, so evolution has chosen hydrophobic amino acids not just because they are hydrophobic but also because they are branched,’ explains Doig. ‘If you want amino acids to go in the core of a protein you make it branched and hydrophobic, if you want it to be on the surface then you make it a straight chain and polar like arginine and glutamic acid.’
Chemical space
Stephen Freeland, an astrobiologist at the University of Maryland in the US, has come up with a method to show that the amino acids adopted by biology were not chosen randomly. He borrowed the idea of chemical space from drug discovery, where molecules are plotted in 3D space to help discover gaps that might reap novel drug molecules. The three parameters investigated by Freeland and his team were size, charge and hydrophobicity. ‘They are not perfect,’ admits Freeland, ‘but as rough proxies of what amino acids do and why they do it, those three are pretty good.’ Hydrophobicity obviously plays a pivotal role in how proteins fold, charge is important in reactions and active sites, and size was just intuitive, Freeland says.
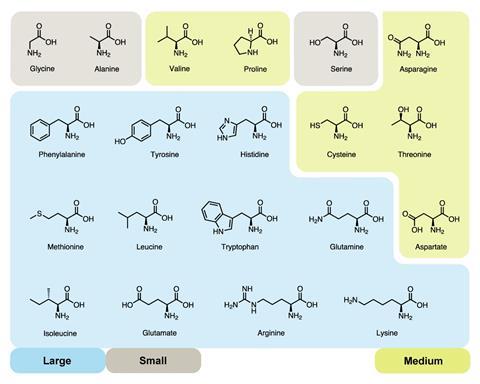
‘We found that the set that is used by biology has a number of surprisingly non-random properties that stand out very clearly,’ says Freeland. The amino acids were widely distributed through their chemical space, but also showed an evenness within that distribution – as if trying to cover as many different property sets as possible.4 ‘What we find with [proteinogenic] amino acids is the moment you build in both of those two factors [hydrophobicity and charge] , just about every test you can throw at them says they are non-random – not only do they cover a good range but they are not clumped to extremes.’
So if this non-random set of amino acids was chosen for good reason, is it possible to create an order in which they were incorporated into biology? ‘There is a consensus now that they didn’t all arrive at once, which to my mind is overwhelming,’ says Freeland. An attempt to come up with a comprehensive order was made by Israeli molecular biophysicist Edward Trifonov, now at the Institute of Evolution at the University of Haifa. Trifonov discovered multiple novel codes in DNA and in the early 2000s turned his attention to amino acids.
Placing the chemically simplest amino acids first might seem obvious, but Trifonov took this further. He looked at multiple criteria, including the energetic cost of their synthesis, the type of transfer-RNA molecules used to transport them and the number of codons (the sequence of three RNA nucleotides that corresponds with a specific amino acid) used in protein synthesis; amino acids with multiple codons are probably older than those with one. He averaged the data and proposed a temporal order starting with alanine and glycine.5
Freeland also looked at how patterns might vary with amino acids assumed to be adopted earlier and later. Using the first 10 alone in chemical space, he found non-random properties in contrast to an examination of all the possible amino acids available on the prebiotic earth (from Miller–Urey or meteorites). Then he added in the complete set of 20. ‘The later ones expand the chemical space of the earlies in ways that maximise range and evenness, and for my money the most interesting single thing is they seem to plug the region of chemical space that was underpopulated, between where the earlies sit and where dimers of the earlies would sit,’ he says. ‘It just makes perfect sense, that this is where you would plug.’
Oxygen expands the code
We certainly know proteins can be made with a much smaller set of amino acids. A Japanese group headed by Satoshi Akanuma at Waseda University recently showed that a 13 amino acid alphabet can create folded, soluble, stable and catalytically active ‘proteins’, albeit not as active or stable as the parent proteins on which they were based.6 So what might have prompted the addition of extra amino acids? According to Moosmann, molecular oxygen forced life to incorporate the last six novel amino acids.
The presumed last six amino acids (histidine, phenylalanine, cysteine, methionine, tryptophan and tyrosine) are all chemically ‘softer’ – they are strongly polarizable and bond covalently. ‘It’s most likely adaptive and not a coincidence or a drift,’ says Moosmann. The idea came to Moosmann during studies on mouse brain tissue (his ‘day job’ involves research into neurodegenerative diseases). He noticed that some amino acids were much more prone to oxidative degradation – those thought to have been adopted later.
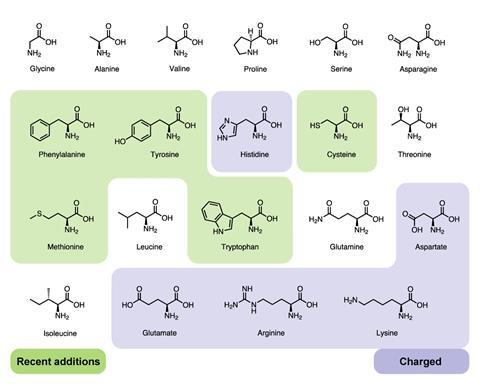
If these amino acids were added to biology for their redox activity he had a hunch that these adaptations were linked to increases in molecular oxygen levels on earth. Oxygen is thought to have become part of the earth’s environment around 2.5 billion years ago in what is known as the ‘great oxidation event’, but Moosmann says that the basic first origin of local low-dose oxygen production is probably older. According to recent research on the evolution of the enzymes involved in photosynthesis, Tanai Cardona at Imperial College London in the UK has suggested the origin of oxygenic photosynthesis to have been 3.6 billion years ago.7
He decided to probe further by looking at the Homo–Lumo gaps for all biological amino acids. The energy gap between the highest occupied molecular orbital and the lowest unoccupied molecular orbital predicts the reactivity of a compound with respect to electron transfer.8 ‘The Homo–Lumo gaps [of all 20 amino acids] had a pattern, just falling at the very point (number 14) when “adaptive” properties came in, and this coincidence is probably not a coincidence!’
The substantially smaller gaps found for the later amino acids suggests their primary function was to undergo redox reactions and Moosmann argues this was needed in an environment where oxygen free-radicals could form, which are particularly destructive to lipids. The ‘softer’ and more redox-active amino acids were capable of protecting cells: ‘These [new amino acid] species could maintain the lipid bilayer integrity in the presence of the rising oxygen concentrations or in the presence of chemical influences which tend to attack or degrade unsaturated fatty acids,’ says Moosmann. ‘For the last three [methionine, tryptophan and tyrosine] there is overwhelming evidence for a response to oxygen.’
One question this then raises is whether our last universal common ancestor contained the full suite of amino acids. A 2016 study identified a set of 355 genes inferred to have been present in the organism that has become known as Luca.9 Moosman says the date for Luca has been placed between 3.7 and 2.9 billion years ago, so it is possible oxygen was available. ‘The consequence of this is indeed that Luca (if it ever existed) had fewer than 20 amino acids.’ He suggests that later genetic code additions could have been distributed laterally to all modern lines: ‘My best guess is that Luca had 17–18 AAs, lacking methionine and tryptophan and perhaps tyrosine.’
Why stop at 20?
Adaptation to an oxygenated world may explain the expansion of the code to 20 amino acids, but why stop there? ‘I would say, look what 20 can do,’ says Freeland. ‘Apparently 20 is good enough for almost every living organism to have adapted to an unimaginable number of habitats over the entire history of life.’
In fact there are at least two additional amino acids used in organisms, although only one of these is found in human proteins – the selenium-containing selenocysteine. It is found in the active sites of 25 human proteins, but is incorporated by a more complex mechanism than normal protein synthesis. ‘This shows that the process had not stopped, it reached a point where incorporating new amino acids is extremely hard,’ says Lluis Ribas, a molecular biologist at the Institute for Research in Biomedicine, Barcelona, Spain. ‘If you want to do it then you need to find very original solutions.’
The limitation is in the recognition of the tRNA
To answer ‘Why 20?’, Riblas has taken a close look at the protein synthesis mechanism – translation. The process is carried out in the cell’s ribosome, a very large complex of RNA and protein molecules. Each amino acid is carried by a bespoke transfer RNA (tRNA) molecule, attached through a hydroxyl group to form an ester. This then reacts with the terminal amino acid of the growing protein chain. The correct amino acid sequence is translated from messenger RNA molecules through Watson–Crick base-pairing with the tRNA molecules. Each tRNA contains a sequences of three bases specific to one of the 20 amino acids – a codon.
Given each amino acid is coded by a sequence of three bases, you might assume there would be 64 possible combinations (of the four possible bases). While three codons are used as instructions to stop protein synthesis, that still leaves 61 – so why stop at 20 unique amino acids? ‘The limitation is in the recognition of the tRNA.’ Ribas says. Each tRNA molecule has a well-defined tertiary structure that is recognized by the enzyme aminoacyl tRNA synthetase, which adds the correct amino acid. From studying tRNA structures, Ribas concluded the problem is finding ways to make new tRNA molecules that could recognise a new amino acid without picking up existing ones.10 The possible structures are limited as they must also fit with the existing protein translation machinery.
‘It’s like if you have a very simple kind of lock where you could only change three or four pins, you come to a point where you wouldn’t be able to make new keys because a new key will open a lock you have already used and that defeats the purpose,’ he explains. The point where nature was unable to create new unique tRNAs that would not be mistaken for others seems to have been at 20 amino acids. In modern biology this allows most amino acids to be coded by more than one codon – the redundancy helping more accurate translation (amino-acid incorporation mistakes are estimated to occur once in 1000 to 10,000 codons).
Expanding the amino acid code
Ribas says his work also has implication for synthetic biologists who are trying to take the genetic code a step further by incorporating unnatural amino acids and perhaps one day improving on nature. In 2011 a team including Harvard synthetic biologist George Church removed one of the three stop codons from E. coli bacteria so it could be replaced with an alternative non-proteinogenic amino acid, and other labs have incorporated such amino acids into proteins.
Evolutionary theory tell us that the set that we have got is a microcosm of what’s possible
But Ribas isn’t sure it will be that this will be a successful strategy for synthetic biologists. ‘If you try to develop a system in vivo for the generation of proteins with unnatural amino acids, it’s not very effective, the efficiency is low, and often you have specificity problems,’ he says. Ribas puts this down to the difficulty in creating new tRNA molecules within the current protein translation machinery. ‘I don’t think there is any way around it, [without] extensive remodelling of the whole machinery;’ although, he adds, this is something currently being done.
Even if it becomes possible, Freeland thinks there will be few advantages. ‘Everything in evolutionary theory tell us that the set that we have got is a microcosm of what’s possible.’ Whether expanding the amino acid repertoire of life will turn out to have useful applications remains to be seen, but there is now plenty of evidence that life’s 20 amino acids were well chosen and not a ’frozen accident’.
But Freeland cautions against an understanding that looks to neatly order chemical evolution. The chances are it was once much messier, with many different types of molecules and mechanisms involved that may have now been replaced. ‘It’s so tempting to work your way up from nothing to something, because that’s what happens when a chemist sits down with a beaker of distilled water and tries to make a reaction happen – but that isn’t what’s happening in the universe, the universe is full of messy chemistry.’
Rachel Brazil is a science writer based in London, UK
References
1 A P Johnson et al, Science, 2008, 322, 404 (DOI: 10.1126/science.1161527)
2 T Koga and H Naraoka, Sci. Rep., 2017, 7, 636 (DOI: 10.1038/s41598-017-00693-9)
3 A Doig, FEBS J., 2016, 284, 1296 (DOI: 10.1111/febs.13982)
4 M A Ilardo and S J Freeland, J. Syst. Chem., 2014, 5, 1 (DOI: 10.1186/1759-2208-5-1)
5 E N Trifonov, J. Biomol. Struct. Dyn., 2004, 22, 1 (DOI: 10.1080/07391102.2004.10506975)
6 R Shibue et al, Sci. Rep., 2018, 8, 1227 (DOI: 10.1038/s41598-018-19561-1)
7 T Cardona, Heliyon, 2018, 4, e00548 (DOI: 10.1016/j.heliyon.2018.e00548)
8 M Granold et al, Proc. Natl Acad. Sci. USA, 2018, 115, 41 (DOI: 10.1073/pnas.1717100115)
9 M C Weiss et al, Nat. Microbiol., 2016, 1, 16116 (DOI: 10.1038/nmicrobiol.2016.116)
10 A Saint-Léger et al, Sci. Adv., 2016, 2, e1501860 (DOI: 10.1126/sciadv.1501860)













2 readers' comments