Researchers measure movement of DNA along motor protein in unprecedented detail
US researchers have adapted a method for DNA sequencing to measure directly and with exquisite sensitivity the movement of a single molecule of DNA as it is drawn through specialised motor proteins. The technique could open the way to much more detailed analysis at the molecular scale of the working of motor proteins, which are involved with myriad DNA- and RNA-processing activities, such as polymerisation, unwinding and cutting-and-pasting.
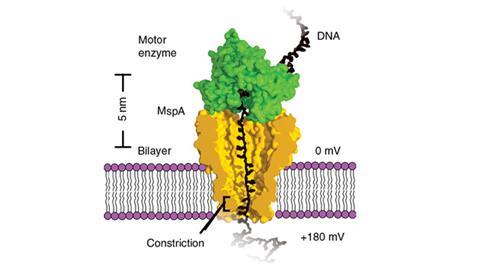
One method of DNA sequencing involves creating a nanometre-sized pore in a membrane that separates two chambers filled with an ionic solution. When an electrostatic field is applied a current flows through the pore and draws a strand of DNA into the pore. The presence of the DNA in the pore alters the flow of the current, whose magnitude can be correlated with the specific nucleotides occupying the pore. If the DNA is gradually drawn through the pore by a motor enzyme, the change in current can be used to infer the sequence of nucleotides.
Jens Gundlach, of the University of Washington, Seattle, and his colleagues realised that with some modification the system could be used not only to sequence DNA but also to observe how the motor protein reels in the DNA strand. The researchers used a mutated bacterial protein to form a tiny constriction in the membrane pore and drew the DNA through the pore with a motor protein called a helicase. They were able to measure the movement of the DNA in steps as short as 40 picometres, or 0.06 of a single nucleotide, a resolution that is unprecedented.
‘We can see how the DNA is being moved through the enzyme,’ says Gundlach. ‘And because we can sequence the DNA at the same time, we can also see how any behaviour, such as pauses during the processing, coincides with particular DNA sequences.’
But Stephen Block of Stanford University, who uses optical tweezer techniques to study the interaction of motor proteins with DNA and RNA at a resolution of around 100pm, wonders whether much is being gained by achieving sub-angstrom resolution. ‘Conventionally, most scientists have therefore concluded that around 100pm, or 1 angstrom, should be ample resolution to detect individual bases. Furthermore, the length of a single chemical bond is just around 100 pm, or 1 angstrom. So the question is, what can one usefully learn about biochemistry below the angstrom? Is there anything new to be learned, or is this just ‘empty resolution’, quite a bit better than is needed to do the job, but not especially useful?’
However, other researchers in the field are impressed. Anatoly Kolomeisky of Rice University in Texas says, ‘This is a great step that provides a new powerful tool in probing complex phenomena in nature,’ while Richard Ebright of Rutgers University believes that the new technique is a ‘transformative, game-changing technology for mechanistic analysis of molecular motors that use nucleic acids as tracks’. Taekjip Ha of the University of Illinois says, ‘I believe this work will open a new era of ultraprecision single molecule analysis, addressing fundamental questions that could not be addressed before due to lack of resolution.’
References
I M Derrington et al., Nat. Biotechnol., 2015, DOI: 10.1038/nbt.3357











No comments yet