PCR primers rapidly reach their binding site with the help of a DNA gliding technique
In the congested intracellular environment, scientists in the Netherlands and the US have found a way for molecules to manoeuvre through the crowds and arrive at their destination faster, increasing the efficiency of lengthy laboratory processes, such as polymerase chain reactions (PCRs).1
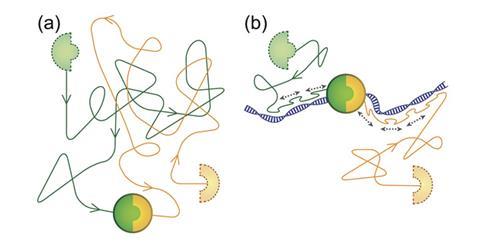
Biomolecules must find and associate with other biomolecules to carry out basic processes within a cell. To do so, many will randomly diffuse within the cell’s three-dimensional space, which is very time-consuming. Some proteins have evolved to glide along DNA and sample positions in a one-dimensional fashion in search of their target, akin to driving down a highway and stopping to sightsee along the way. This has the potential to speed up these processes.
Antoine van Oijen at the University of Groningen and colleagues have given proteins a chance to ride the DNA highway that would otherwise be incapable of doing so to see if this would enhance their association speed with target proteins. As proof of concept, he took advantage of the well-known velcro-like properties of biotin and streptavidin, which form tight molecular interactions with one another after diffusing through a solution. Instead, an 11 amino acid long DNA interacting peptide, pVIc,2 from adenovirus was glued to these biomolecules. By adding these components in solution along with stretches of DNA, the peptides bound non-specifically to DNA and travelled along to find their partner, decreasing association times by up to 20-fold.
Reducing dimensionality to accelerate biotin and streptavidin binding depended on the DNA base pair concentration rather than DNA length. Michael Schlierf, an expert in protein-DNA interactions at the Technical University of Dresden, Germany, explains: ‘In other words, not the length of a single road, but the length of all roads is important.’
Van Oijen then applied this concept to PCR, a known laborious process to amplify DNA. A key step in this procedure is when oligonucleotide primers bind to specific DNA sequences, initiating replication. Primers coupled with the adenoviral peptide bound non-specifically to DNA, searching and reaching their intended sequence more efficiently. The result was not disappointing. The reaction time decreased by 15 to 27%, meaning it generated the amplified DNA product 5 to 7 cycles earlier than in a typical reaction.
Hagan Bayley, a chemical biologist at Oxford University, UK, praises this development and hopes it will lead to greater clarity in the field. ‘They have revived the issue of one-dimensional diffusion, but with a practical twist. The ability to speed up and redirect reactions by this means has real utility.’ Sy Redding, a chemical physicist at the University of California, San Francisco, US, offers a slightly different perspective: ‘The general hustle and bustle of molecules working and running around on DNA may not be as much of a traffic jam as it seems. In fact it may be that the traffic is precisely what allows the cell to ensure proteins get to where they are supposed to be on time.’
Having now relocated to the University of Wollongong in Australia, Van Oijen tells Chemistry World he is excited to build on these results and cautions there is still room for improvement. ‘The PCR industry has already evolved to become quite efficient. Therefore, the obvious next step is to provide PCR reactions with more specificity in complex reaction mixtures and protocols.’
References
1 A Turkin et al, Chem. Sci., 2015, DOI: 10.1039/c5sc03063c (This article is open access.)
2 P C Blainey et al, J. Biol. Chem., 2013, DOI: 10.1074/jbc.M112.407460









No comments yet