With enough understanding and computing power, it should be possible, but will it happen?
I started working in drug discovery exactly 30 years ago, and one of the newest additions to the hallway where I worked was a room marked CADD: Computer-Assisted Drug Discovery. This was stocked with things like Evans and Sutherland work stations, which now look as if they should have a museum guide next to them explaining what they might have once been used for, but at the time I was impressed. The older generation of medicinal chemists across the hall was less so: they had put a sign up over their own door reading BADD, for Brain-Assisted Drug Discovery, and there matters stood.
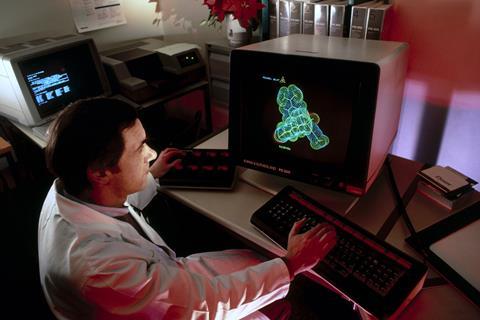
There was a lot of optimism on the CADD side of things at the time, optimism that now seems touchingly misplaced when you consider the hardware and software with which these visions were to be realised. It’s very hard to keep one’s recollections from being tinged with hindsight, but I can claim both enthusiasm and wariness. The former, because I have always been a great lover of new technology, and the latter, because I had had some disconcerting experiences with early molecular mechanics software in graduate school (watching the program cheerfully minimise onto weird or obviously incorrect structures). But the hope was that we would soon be able to compute our way to drug structures, or at the very least rank-order our own ideas to save time. Thus would the CADD and BADD worldviews be reconciled.
Well, the years have gone by, and our computers are wildly more capable, but we’re still not exactly cranking out wonder drug structures at the push of a button. Problems that seemed a few years away from being solved back then now seem… still a few years away from being solved. Hydrogen bonds, for example, are irreplaceably crucial currency in evaluating biomolecules’ behaviour, but they’re still not easy to model accurately. Water molecules, which are tiny hydrogen-bonding machines, are likewise crucial to get right, and that’s proven very difficult to do well. When some ligand displaces a single water molecule from a protein’s binding site, does that help or hurt the enthalpic and entropic terms in the free energy equation? The answer can only be ‘Yes and no and maybe and maybe not’, because all those combinations are feasible depending on the situation. The amounts of energy involved in moving even a single water molecule in or out of position can be enough to take a compound from potent binding at nanomolar concentrations to completely inactive, and back again.
We also have a greater appreciation for molecular dynamics – the realisation that we’re not just modeling a static scene in isolation, but rather looking at the result of a whole set of motions and trajectories as all those energies balance themselves out. This of course requires even greater understanding (and greater computational resources). But if you’re worried about the accuracy of one of those static pictures, how do you feel about frame after frame of them going past in a molecular dynamics simulation? Exhilarated, or even more wary than before? An accurate look at dynamic processes could be like having magic powers, but an inaccurate one gives you new chances to be deceived 30 times a second.

It’s always been true, though, that there don’t seem to be any inherent reasons why such simulations can’t work. The same goes for all those other tasks we would like computational chemistry to do for us. Realising them calls for deeper understanding, better models, more accurate estimations – enough of all these and there’s really no reason why you shouldn’t be able to calculate your way to a reasonable small-molecule ligand. It’s just that getting to ‘enough’ on these has been a much slower process than the optimists in the crowd hoped for back in the 1980s. One can be a long-term optimist and a short-term pessimist in these matters, willing to believe that these are solvable problems but holding on to your wallet when someone claims to have solved them right now.
















No comments yet