AlphaFold, an AI program that has previously demonstrated that it can predict protein structure from an amino acid sequence, has been paired with two other AI routines to afford an end-to-end AI drug discovery process even when a protein structure is not known.1 This combination of machine learning processes was able to predict a novel drug-like small molecule against a new target for liver cancer, demonstrating how AI can design bespoke therapeutics rapidly and accurately.
‘For many targets that are implicated in disease there are no known structures. Using a structure provides advantages to that design process that other methods cannot compare with,’ explains Petrina Kamya, head of AI platforms at Insilico Medicine, the company which developed the technique with input from Alán Aspuru-Guzik of the University of Toronto, Canada, and Nobel laureate Michael Levitt of Stanford University in the US.
The structure of a target protein is normally obtained from experimental methods such as x-ray crystallography. However, there can be difficulties in obtaining these structures and for entirely novel targets acquiring an x-ray structure may be time consuming.
‘Cue AlphaFold, a computer program that promises to solve this age-old protein folding problem by claiming to have generated the structures of all proteins in the human genome,’ says Kamya. To show the benefit of using AlphaFold to generate a target protein structure, the team at Insilico Medicine paired AlphaFold with two AI programs they have developed; PandaOmics and Chemistry42.
PandaOmics can sift through, and interpret, a wide range of omics data. Omics data is typically obtained from high-throughput biochemical assays and this information is used alongside text mining of the scientific literature and grant descriptions in the hunt for possible therapeutic targets. ‘PandaOmics is a target discovery engine that provides insights into disease–target relationships that may have been previously overlooked, with plenty of evidence to back up the connection,’ explains Kamya.
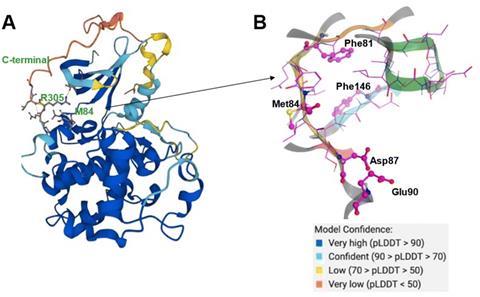
Aiming towards a small molecule that could be used against hepatocellular carcinoma, a common liver cancer, the team at InSilico harnessed the interpretive power of PandaOmics to uncover the protein CDK20 as a reasonable target for treatment. CDK20 is overexpressed in tumour cell lines and a reduction in this protein in lung cancer cells is implicated with reduced cell proliferation and an increased sensitivity to radiochemical treatments. While no protein structure is known from experiments, AlphaFold was able to suggest a likely structure of CDK20.
30 days from target to hit
Next, the team paired the AlphaFold predicted structure with the generative AI Chemistry42, which suggested binding sites for a small molecule inhibitor of CDK20. By analysing the AlphaFold-predicted protein structure, Chemistry42 suggested a shallow binding pocket of 150Å3. Further investigation of this pocket via the AI’s generative routine saw it knit together plausible chemical structures, moieties, and functional groups to suggest a range of ligands. These ligands were designed around the 3D shape of the binding pocket within CDK20, the volume available and the spatial arrangements of key atoms and protein residues. The AI suggested 54 potential inhibitors, of which the team synthesised seven. This process took only 30 days from target selection to the first hit, and subsequent refinement from Chemistry42 discovered even more potent hit molecules that demonstrated anticancer activity in experimental testing.
‘Compared to the timelines and investments required in drug discovery campaigns, this fully AI-driven approach has demonstrated huge advantages and shows a paradigm shift in drug discovery entirely driven by AI predictions and de novo generation of chemical structures,’ remarks Pablo Carbonell, a computational biologist at the University of Valencia in Spain.
For the team at Insilico Medicine, one of the next goals is incorporating end-to-end AI drug discovery into robotic labs, such as their new facility in Suzhou, China. Aspuru-Guzik explains that self-running labs could combine AI-led drug discovery with automated reactions and drug formulations2 suggested by machine learning for an accelerated process.
‘I think the next natural evolution of this is public open drug discovery,’ adds Aspuru-Guzik. ‘There should be federally funded institutions to work on neglected diseases with a non-profit nature. As the tools become cheaper and more democratic, why not create a federal agency for this?’
References
1 F Ren et al, Chem. Sci., 2023, DOI: 10.1039/d2sc05709c
2 P Bannigan et al, Nature Commun., 2023, DOI: 10.1038/s41467-022-35343-w










1 Reader's comment