Measuring floppiness might help untangle the structure of molecular knots, catenanes and other mechanically interlocked molecules, a team of chemists has proposed. The floppiness factor – determined by mass spectrometry (MS) – helps distinguish between molecules interlocked in different ways even if they have the same mass.
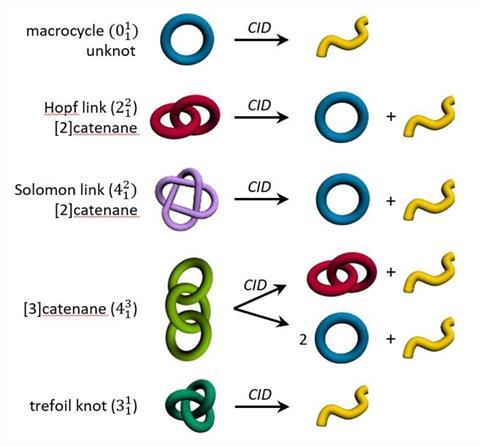
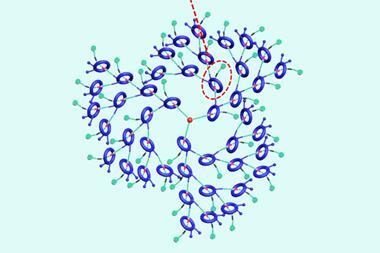
Mechanically interlocked molecules – like trefoil knots – are often tough to characterise. Nuclear magnetic resonance spectra are crowded with tens of overlapping signals that tell chemists little about the molecule’s topology. In mass spectrometry, catenanes often give the same signals as the rings and the linear fragments that make up the molecule. Moreover, there’s no way to distinguish between different architectures, like simply interlocked rings – called a Hopf link – or doubly interlaced Solomon links.
An international team of chemists has now changed this. ‘We started thinking about the difference between Hopf link and Solomon link, and saw that they differ in how strongly the two rings are intertwined with one another,’ says the study’s first author Anneli Kruve from Stockholm University, Sweden, who conducted the work as a visiting researcher at the Free University Berlin, Germany.
Through mass spectrometry experiments, Kruve and colleagues determine what they call a floppiness factor – a measurement of how entangled a molecule’s individual parts are. A Hopf link is floppier than a Solomon link, and a three-ring linear chain floppier than a trefoil knot.
The floppiness factor compares a linear fragment’s behaviour with the parent catenane. To calculate it, the compound is first run through collision-induced dissociation, which breaks up the catenane into individual rings and linear fragments. This is followed by ion mobility mass spectrometry, in which the fragments are separated based on the interactions with a buffer gas before reaching the mass analyser.
‘In the synthesis of these compounds, one very important bit of information is how do they interlock,’ says Ho Yu Au-Yeung, who studies catenanes at the University of Hong Kong. ‘It’s a very challenging task, and the more we go to more complex interlocked structures, the more difficult it gets to characterise them. That’s why I think such additional information on the compactness by this new MS experiment is really important.’
The only limitation, Au-Yeung says, is the need for an ion mobility mass spectrometer, which is not as widely used as ordinary MS instruments. He hopes the method will be developed even further, to not only compare topological isomers, but determine an absolute compactness measure for catenane characterisation.
Kruve says there’s room for adapting the method to other interlocked molecules, like rotaxanes. ‘In general, ion mobility could be very useful to differentiate, for example, between a rotaxane and just a side-on complex.’
References
A Kruve et al, Angew. Chem., Int. Ed., 2019, DOI: 10.1002/anie.201904541












No comments yet