The longstanding goal of designing RNA-targeting drugs has been given a boost thanks to a technique that identifies drug-like small molecules which selectively bind to disease-causing RNA. The US team demonstrated the approach by designing an antiviral drug lead for hepatitis C based on its RNA sequence and suggest the technique could help combat new viral strains that emerge seasonally such as flu.
Traditionally, drugs are designed to bind to the folds of proteins to block or disrupt activity that is responsible for illness. However, RNAs are an attractive therapeutic target because they are responsible for encoding protein production. Moreover, the bulk of RNAs in the human genome are noncoding and many are known to be associated with disease, including genetic disorders and cancers. But RNA has been an elusive target, largely because knowledge is lacking, both on which RNA folds accept binding of small molecules and which small molecules bind to those RNA folds.
‘This information is generally not known and having it would allow one to advance a gene-to-RNA-to-medicine pipeline that we have been trying to do,’ says Matthew Disney, whose lab at the Scripps Research Institute has been working on the problem of targeting RNA for over a decade. ‘People did not think that RNA could be selectively targeted with small molecules and I think we have shown some examples where indeed one can.’
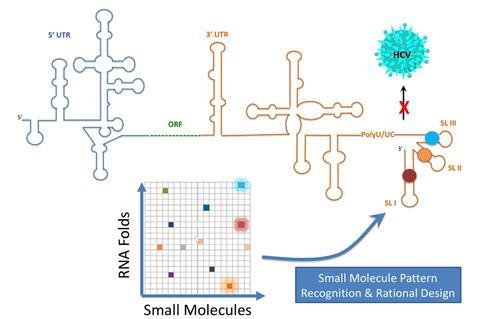
Building on their recent work, including a screening platform called Inforna that has identified small molecules that bind to disease-associated RNAs, Disney and colleagues have gathered 10-times more information about interactions between RNA folds and small molecules than was previously known. Armed with this information the team expanded the concept to viral RNA by identifying sites in the hepatitis C genome that small molecules can bind to.
Molecular mapping
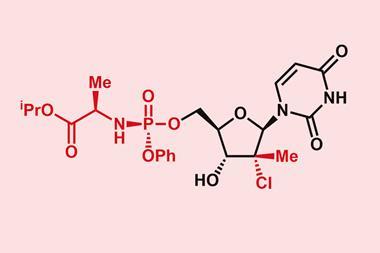
To gather the required binding information, the researchers screened a library of 30,000 small molecules to find almost 2000 compounds that were drug-like. These were then screened for binding against libraries of RNA folds. This identified which of the 52 million possible interactions might be a fruitful avenue to explore for drug candidates. The team found that 239 compounds bound RNA and used the results to make maps of small molecule RNA recognition patterns. By comparing the interaction maps with the hepatitis C virus genome, it was possible to identify viral RNA sequences that had partnering binding sites for specific small molecules, which led the team to identify a small molecule that inhibited viral replication.
‘It shows that one can rapidly take chemical libraries that are everywhere – from academia to industry – to get compounds that bind RNA to affect biology for a favourable therapeutic benefit,’ says Disney. ‘One of the advantages of this approach – in principle – is that as a seasonal flu or seasonal virus emerges, one could identify the sequence and then design small molecules rapidly to target the folds from the sequence.’
‘Currently, there is considerable debate about whether RNA is “druggable” and whether the compounds that target RNA will be similar structurally to those that target proteins,’ comments Steven Zimmerman, who investigates RNA-targeted small molecules at the University of Illinois, US. ‘This paper adds support, suggesting that the structures effective for RNA will be analogous to traditional drugs. Interestingly, some of the most effective compounds for targeting RNA violate one or more of Lipinski’s rules [a rule of thumb used to evaluate how drug-like a molecule is], suggesting that our view of what constitutes a drug needs to be expanded.’
‘We want to scale this even further and make the design of small molecules targeting RNA very routine and easy. This could change a paradigm in drug discovery where RNA is an important drug target for small molecules,’ says Disney. ‘These leads would start someone on the long and winding road to getting an actual medicine to a patient.’
References
J L Childs-Disney et al, Chem, 2018, 4, 1 (DOI: 10.1016/j.chempr.2018.08.003)








No comments yet