Scientists have developed a method for purifying large numbers of peptides in a parallel fashion. It uses a cleavable linker system and releases unprotected peptides. The team hope to develop the concept into a routine tool that can keep pace with the increasingly high output of parallel peptide synthesis.
Recent developments in parallel solid-phase peptide synthesis can yield hundreds of peptides per synthesis. Traditional approaches to remove byproducts from the resulting peptide sample rely on high-performance liquid chromatography (HPLC). However, the efficiency of HPLC decreases as peptide length increases with greater numbers of incomplete reactions to match, resulting in more impurities and associated separation challenges. As HPLC is a linear process, even the most advanced purification set-up can take over 15 minutes per peptide. Multiply that for tens of peptides and you encounter a severe bottleneck.
‘Our approach was focussed on preventing side reactions by creating a reductively cleavable linker to increase purity,’ explains Robert Zitterbart from peptide-therapeutics company Belyntic in Germany. Zitterbart and his colleagues have discovered that adding a brominated para-azido benzyl moiety linker to the peptides as the last step during peptide synthesis held the key to traceless purification.
The linker immobilises the target peptide onto a solid support so that all non-tagged impurities such as byproducts or other fragments lacking the linker can be removed by washing. Next, the linker is primed for cleavage by reducing its aryl azide to an aniline, and the reducing agent is washed out using water. In the last step, acid catalysed 1,6-elimination breaks the linker under mild conditions. This is an advantage over using basic conditions to detach the linker, as they can disrupt peptide integrity.
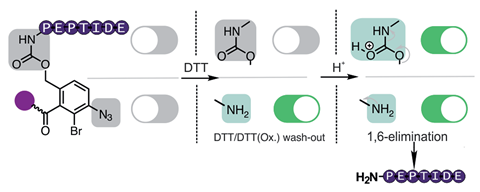
‘The most rewarding aspect of this challenge was fine-tuning the electronic properties of the linker molecule to give the release mechanism,’ explains Zitterbart. ‘We ended up choosing halogenation, with bromine on the para-azido-benzyl core, because it is relatively easy to introduce and has a high electronic withdrawing effect.’
As a proof-of-concept, the team used their technique to purify a personalised cancer treatment consisting of 20 neoantigen peptides. 14 of those peptides achieved a final purity of 90% within a six hour timeframe, pointing at the technique’s potential to speed up therapeutic manufacturing. ‘It’s like the difference between a dishwasher, washing lots of items in one go, and with washing each item of dirty cutlery by hand, and changing the water several times in between,’ says Zitterbart.
‘This upgraded linker is an improvement on the previous Peptide easy clean technology – which suffered from base-mediated side reactions in some sequences – because it can be mildly activated by reduction of an aryl azide. The use of classic physical organic parameters in the linker design and optimisation – to increase stability to trifluoroacetic acid cleavage and avoid bromination byproducts – is elegant and intellectually satisfying,’ comments Jenn Stockdill, whose research at Wayne State University in the US explores how to increase the synthetic accessibility of peptide and protein-derived therapies.
But the approach is not without its drawbacks. Sometimes impurities that form in the full-length peptide chain during solid phase peptide synthesis are not always removed. However, the team says their approach can act as a general purification step to get a significantly lower overall proportion of impurities, at which point HPLC purification becomes viable once more. Zitterbart says it ‘opens the door to purifying more challenging peptides such as hydrophobic peptides that would otherwise stick to the phase columns, or peptides with multiple truncations. It also uses significantly lower amounts of high purity grade organic solvents than typical HPLC processes, which could help drive peptide manufacturing towards a more sustainable future.’
Stockdill says testing the technique on personalised peptide vaccine libraries ‘was an exciting application that knocked my socks off. Three thumbs-up! I can’t wait to have my students use this method to purify our peptides. They’re going to be so excited to not wake up at 4am to get on the HPLC.’
Paragraph eight of this story was updated on 17 February 2021 to clarify the drawbacks of the technique









No comments yet