Trapping amides inside a molecular cage small enough to twist them out of shape makes them up to 14 times more reactive, chemists in Japan have discovered. This mechanical twisting method is inspired by the way enzymes break peptide bonds and could eventually lead to catalysts that work under ambient conditions.
Amides are less reactive than many other functional groups, such as esters, because of resonance stabilisation between the carbonyl’s π* orbital and the nitrogen’s lone pair. This might be one of the reasons amides – in the form of peptides – are the bread and butter of biochemistry. Strongly alkaline or acidic conditions are needed to hydrolyse them. In water at room temperature, a peptide bonds’ hydrolysis half-life is at least 350 years.
However, the resonance that makes amines inert can be broken by forcing the bonds out of planar alignment. Chemists have made use of this mechanism by creating amides with twisted cyclic structures or by attaching bulky substituents that bend the amide when coordinated to a metal. ‘In our work the method for twisting is different, it’s non-covalent,’ explains Hiroki Takezawa from the University of Tokyo, who led the study alongside Makoto Fujita.
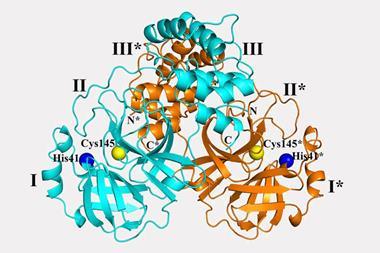
Takezawa and colleagues found that amides twist into a reactive shape when they become trapped in a molecular cage. A similar mechanism has been proposed for peptide bond-breaking enzymes, in which a reactive pocket contorts the amide into a bent conformation.
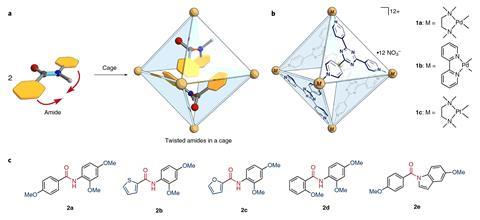
The octahedral cages consist of six palladium or platinum atoms connected by four electron-deficient ligands. ‘The containers for the reactions are incredibly easy to make,’ comments supramolecular chemist Bruce Gibb from Tulane University in the US. ‘And that always plays into their utility.’
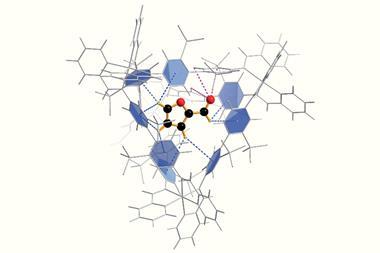
Stirring the cage compound with an electron-rich amide in 100°C water for an hour creates the inclusion complex. For some substrates, like N-(2,4-dimethoxyphenyl)-4-methoxybenzamide, two molecules can fit inside the cage, both adopting a twisted shape. This speeds up the amide’s hydrolysis by a factor of five.
For N-(2,4-dimethoxyphenyl)thiophene-2-carboxamide the team needed to use a phenanthrene modulator. This compound sits inside the cage alongside a single amide molecule, ensuring that it twists into its reactive conformation. In this co-inclusion complex the amide’s hydrolysis rate is enhanced 14-fold.
How exactly different amides behave inside the cage isn’t something his team could predict, Takezawa says. This shows that ‘we’re a long way from being able to control and understand how molecules in confinement react,’ says Gibb. ‘Although this paper does not demonstrate full control on the confirmation of the encapsulated amide, it does importantly demonstrate a proof of principle that by encapsulation we can control conformation and hence reactivity.’
Takezawa points out that the biggest hurdle to practical applications is product inhibition – once the amide has reacted, the products are stuck inside the cage. ‘I think the catalytic reaction can be realised, maybe with a different solvent system or by correctly designing the starting materials,’ he suggests.
Now, his team not only wants to test the caging strategy on more and different amides, but also see if they can influence the reactivity of alkynes and alkenes by confining them.
References
H Takezawa, K Shitozawa and M Fujita, Nat. Chem., 2020, DOI: 10.1038/s41557-020-0455-y







No comments yet