Stripped down bacterium has fewest genes of any replicating organism and will help scientists understand gene function
US researchers led by genomics pioneer Craig Venter and Clyde Hutchison have created a bacterium with a minimal genome containing just 473 genes that are essential for it to survive and replicate. The stripped down genome is smaller than anything known to exist in nature and could enhance our understanding of life and how to tweak it for novel applications.
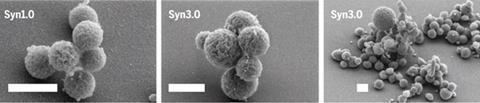
Venter’s team at the J Craig Venter Institute in La Jolla, California, made the first synthetic cell back in 2010, dubbed JCVI-Syn1.0. To create it, they built an artificial replica – but containing some genetic ‘watermarks’ – of the genome belonging to the microbe Mycoplasma mycoides via chemical synthesis. This was then inserted into an existing bacterial cell of another Mycoplasma species which had its DNA removed. In 2014, Jef Boeke’s team at Harvard University went on to create a fully synthetic yeast chromosome from scratch.
Now the team have weeded out the genes of the Mycoplasma mycoides genome that are unnecessary for the bacterium to survive in lab conditions. The result was a minimal cell, called JCVI-Syn3.0, comprising 473 genes, which is almost half the number of genes in the naturally occurring species. It’s also 52 fewer genes than the bacterium with the smallest known genome which belongs to the species Mycoplasma genitalium.
‘There is still not a single self-replicating cell in which we understand the function of every one of its genes,’ explains Dan Gibson, who co-authored the study. ‘The generation of a minimal cell is a major step toward understanding how cells work at the most fundamental level. It is also useful as a chassis for industrial applications or at least as a launch pad for making more complex organisms with extraordinary properties.’
To create their minimal genome, the team developed a protocol to gather data on which genes were dispensable. This involved inserting genes with transposons – foreign DNA sequences that can jump around the genome – to disrupt gene function and determine which were crucial for the cell to function. Combined with computational analysis and the generation of combinatorial libraries of reduced genomes, the team used the transposon data to design, build, and test thousands of possible genome constructs until they whittled it down to the bare minimum.
Even with its minimal genome, the biological function of 149 – almost a third – of the genes in JCVI-Syn3.0 remain unknown. But Gibson says that the relative simplicity of JCVI-Syn3.0 should make these functions easier to discover. ‘The next phases of this project will be to understand what every gene does.’
‘It represents another landmark study in our ability to redesign natural systems at the genetic level using synthetic genes and design constraints,’ comments Paul Freemont, co-director of the Imperial College Synthetic Biology Hub in London, UK. ‘However the future applications of such a technology are less clear given the new genome editing techniques that are now readily available and perhaps more accessible such as Crispr and Cas,’ he adds.
References
C A Hutchison III et al, Science, 2016, 351, 6280 (DOI: 10.1126/science.aad6253)









No comments yet