Deaths in a clinical trial might have been caused by a covalently binding drug that wasn’t selective enough for its target
The case of BIA 10-2474 is a terrifying one. It’s an experimental inhibitor of the enzyme fatty acid amide hydrolase (FAAH), a protein target with a number of potential uses in pain relief, anxiety, and several other conditions. Many readers will remember, though, that the compound was given to human volunteers in early 2016 with catastrophic results. One person died, and several endured severe neurological injuries, even though no such effects had been noted in animal studies. Since then, the question has been how this happened, and how it could have been allowed to happen at all.

A group led by Mario van der Stelt at Leiden University in the Netherlands has some possible answers. BIA 10-2474 was known to inhibit FAAH by a covalent mechanism – it forms a new bond to an active-site amino acid and thus shuts down the enzyme. This can be a very effective way to target a protein, and a number of widely used drugs (such as aspirin and all the penicillin/cephalosporin antibiotics) work this way. But at the same time, covalency has risks. A compound that can react with one protein can almost certainly react with others, leading to off-target shutdown of other biochemical pathways. The resulting altered proteins can also, under some conditions, set off allergic immune responses when they suddenly become recognised as foreign (which is the mechanism behind poison ivy’s effects on the skin). Anyone developing a covalent drug has to be aware of these complications.
And these are most likely the cause of this compound’s terrible effects in the clinic, since several other possibilities have by now been ruled out. The actual drug samples appear to have been uncontaminated, and FAAH inhibition itself is unlikely to be the problem, since other inhibitors have gone into human trials without incident. This study seems to be the first that tries to profile the range of proteins that BIA 10-2474 can react with. I should note that if Bial – the Portuguese company developing the compound – has done such experiments, they have not come to light.
This activity-based protein profiling is one of the key techniques in chemical biology. The idea is that you modify the initial compound by adding a small structure to it such as an alkyne, and verify that such analogues maintain the activity of the original drug. The alkyne serves as a reporter: after the compound has been exposed to mixtures of proteins or living cells, and has formed covalent bonds to its targets, a tag can be attached to all the modified alkyne-containing proteins using an azide–alkyne cycloaddition ‘click’ reaction. This tag (for example a fluorescent group or biotin) can then identify the proteins involved through gel assays, mass spectrometry, protein sequencing, and so on.
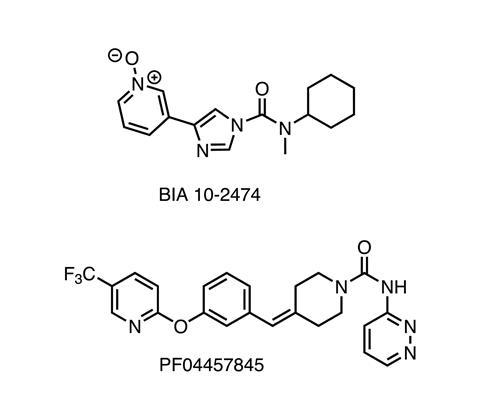
Applying this to BIA 10-2474 and comparing it to an earlier FAAH inhibitor (PF-04457845, also a covalent drug, but without the terrible side effects) showed that Bial’s compound did indeed bind to and inhibit a whole list of other enzymes. Many of these are lipid hydrolases that are strongly expressed in brain tissue, and moreover, their mouse forms are much less reactive with BIA 10-2474 than their human homologues. Taken together, these findings strongly suggest where to start looking for the cause of the clinical disaster and how it might have occurred.
Well, they do at the level of biochemistry, anyway. At the higher level of decision-making and risk management, there were clearly even more severe problems. It is not at all clear that the company realised the extent of the possible risks of developing a covalent inhibitor of an enzyme in the central nervous system. They may have believed that their compound would behave similarly to PF-04457845, without realising the other enzymes that it was targeting (and the differences that could make rodent toxicity results unsuitable for predicting problems in human subjects). But nothing can be assumed in drug development, and everything has to be checked. Attempts to bypass these can result in trips to the emergency room – or the cemetery. Such are the destinations of shortcuts.
References
A C M van Esbroeck et al, Science, 2017, 356, 1084 (DOI: 10.1126/science.aaf7497)

















No comments yet