Where are the next generation of antibiotics going to come from? Clare Sansom looks at the pipeline
The threat of antibiotic resistance is big news. Few people will have failed to notice headlines in recent years warning of a potential ‘antibiotic apocalypse’ that would render most modern medical techniques impossible, or at least fraught with danger. And this is not just media hype: it seems that the more people know about this issue, the more worrying they find it. Already, drug-resistant bacterial infections kill 700,000 people every year, over 90% of them in low and middle-income countries, and authoritative sources suggest that this figure may rise to 10 million by 2050, more than cancer kills today.
Antibiotic resistance has been known about almost since the dawn of the antibiotic era. As early as 1945, Alexander Fleming, the discoverer of penicillin, recognised that the drug’s over-use could lead to bacteria developing resistance. ‘In such cases, the thoughtless person playing with penicillin is morally responsible for the death of the man who finally succumbs to infection’ he told the New York Times. Sally Davies, England’s Chief Medical Officer since 2010, is one of the issue’s most powerful current advocates. Her voice was one of those that called on UK Prime Minister David Cameron to commission a wide-ranging review of the issue in 2014. That review, chaired by the economist Jim O’Neill and published two years later, set out 10 specific steps to reduce the threat. Many countries now have ambitious plans for action, and the issue has been discussed in the United Nations at the highest level. Yet Davies is still not satisfied: ‘We need to up the ante,’ she says.
Several organisations have drawn up lists of ‘priority pathogens’ – bacterial species in which the problem of resistance is most severe and the need for new drugs particularly acute. The Infectious Disease Society of America’s five priorities of Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter species, are known acronymically as the Eskape pathogens. The World Health Organisation’s list of 10 is divided into three groups according to the severity of the threat they pose: the most urgent, critical group includes all the Eskape pathogens apart from Staphylococcus. It is instructive that almost all the bugs on these lists are the so-called Gram negative bacteria with a double-layered cell wall: that wall is hard for small molecules to penetrate so it is not surprising that these are the hardest bacteria to design drugs to target.
Under-resourced and under-funded
A successful antibiotic must kill bacteria – or at least prevent them from multiplying – while causing minimal damage to the human body, so it must attack a molecular target or mechanism not found in human cells. These are either wholly unique to bacteria, such as their cell walls, or differ enough in structure and mechanism from mammalian ones to be inhibited selectively. The largest class of antibiotics, the beta-lactams, which are named after their strained ring structure, fall into the first category: they disrupt bacterial cell wall synthesis by inhibiting the formation of the peptidoglycan layer of that cell wall. Drugs that bind to bacterial ribosomes and target the synthesis of their proteins fall into the second.
Antibiotic resistance is far older than man-made or even repurposed antibiotics. Bacteria have been evolving ways to protect themselves from harmful substances for billions of years, and this resistance can spread extremely rapidly, mainly through so-called ‘horizontal’ gene transfer between organisms. The commonest three mechanisms involve breaking down the antibiotic molecules themselves, modifying target proteins so that the drugs no longer bind or ejecting the drugs from the cells before they can take effect.
Resistance to the antibiotics of the past and present would matter a lot less if many new ones were coming down the pipeline. The reality, however, is very different: only a handful of the new chemical entities (NCEs) registered by the FDA and EMA in recent years have been antibiotics, and these have all been what the industry calls ‘me-too’ drugs: existing compounds modified with small tweaks that might, for example, reduce side effects or improve their performance against resistant strains. Antibiotics with novel targets, or even drugs that bind to the same target but have a different mechanism of action, will be harder for bugs to beat. Historically, of course, these have been few and far between, but recent decades have seen none at all.
It is self-evident that antibiotic discovery needs significant investment and a large, well-trained scientific workforce. For some years, however, big pharma companies have been withdrawing from the antibiotic market in search of the larger and more certain profits found in drugs for chronic disease. A survey of antibiotic research and development by PricewaterhouseCoopers indicated that fewer than 150 industrial scientists were working in this field in the UK in 2016. At least 40 of these jobs have been lost since then when Redx Pharma closed its anti-infectives unit and AstraZeneca sold its antibiotics research business to Pfizer. In contrast, just one charity, Cancer Research UK, supports about 4,000 researchers. Figures from other countries are roughly comparable, and something similar can be seen in clinical practice. In the US, infectious disease physicians are less well-paid than those in almost any other specialty and it is increasingly hard to fill vacancies. It is not surprising that one of the O’Neill review’s headline recommendations is to ‘improve the numbers, pay and recognition of people working in infectious disease’.
Working together
Small biotech companies, often with academic partners, are now stepping into the gap left by the pharmaceutical behemoths, and, at last, substantial funding is becoming available to support innovative development programmes. The CARB-X initiative, set up in mid-2016 with funding from the US Department of Health and the UK’s Wellcome Trust, expects to invest almost half a billion dollars (around £350 million) over five years to accelerate the development of at least 20 novel products – antibiotics, diagnostics and vaccines – targeting the most critical resistant pathogens. ‘It is early days, but we have already invested in 24 projects involving 10 novel molecular targets,’ says Kevin Outterson, Executive Director of CARB-X. ‘I was born in the year when the last new approved antibiotic class for treating Gram negative organisms was discovered [1962], so if just one of our projects makes it all the way to approval it would be the biggest step-change in antibiotic research in my lifetime.’
The UK R&D Centre for Antimicrobial Resistance (AMR Centre), based at Alderley Park in Cheshire on a site previously occupied by Astra Zeneca, was set up in 2016 with a similar aim to CARB-X but with rather smaller sums to disperse. It plans to invest $6 million in its first year, with a focus on the WHO’s ‘critical’ pathogens. ‘We aim where possible to invest in programs alongside CARB-X and we collaborate extensively, but there are differences between our portfolios,’ says Peter Jackson, executive director of the AMR Centre. ‘Unlike CARB-X, the AMR Centre has labs on site, so we can offer smaller companies the facilities they might need for, for example, drug metabolism and toxicity testing, formulation or in vivo studies. We can also get involved a bit later in the drug discovery pathway, from lead optimisation up to Phase 2 clinical trials.’
These and similar initiatives are helping to increase the number of scientists involved in antimicrobial research, as well as contributing to two further research-based recommendations of the O’Neill Review: ‘[to establish] a Global Innovation Fund for early-stage and non-commercial research’ and ‘[to provide] better incentives to promote investment [in] new drugs and improve existing ones’. Encouragingly, the antibiotic pipeline is at last beginning to widen, with more novel molecules coming through the early stages.
CARB-X and the AMR Centre also provide opportunities for the companies they fund to network and to share information and resources. Such inter-company collaboration is becoming more common particularly in therapeutic areas such as infectious disease where expected financial rewards are small. The ability to share data provides a further boost to collaboration, and antibiotic research can now benefit from a new digital platform for data sharing: The Pew Charitable Trusts’ Shared Platform for Antibiotic Research and Knowledge (Spark). This uses technology by Collaborative Drug Discovery, an international software company based in California, US. ‘Our platform, CDD-Vault, provides a complete, secure informatics interface for drug discovery, where companies and their academic collaborators can share data exactly how and with whom they like,’ says Mariana Vaschetto of the company’s Cambridge office in the UK.
A long history
Few academic institutions have had as long and as productive a history of antibiotic research as the UK’s University of Oxford. Howard Florey, who shared the 1945 Nobel prize for medicine with Fleming and Ernst Chain, held a chair in pathology there, and the first patient to be treated with penicillin – a policeman with an infected wound on his face – was treated at the city’s Radcliffe Infirmary in 1941. The policeman initially responded well but died when supplies of penicillin ran out. Penicillin chemistry has continued at Oxford under Edward Abraham, Jack Baldwin and now Chris Schofield. It is often forgotten that Dorothy Hodgkin, also a professor at Oxford, won her Nobel prize in 1964 not for her best-known molecular structure, insulin (which was not completely deciphered until 1969) but for two simpler ones: vitamin B12 and penicillin.
Current work in Schofield’s group focuses on understanding the structure and mechanism of beta-lactamase enzymes that have evolved in many bacteria to combat beta-lactam antibiotics. These all break open the antibiotics’ ring structure to render them useless, but they use two different mechanisms to do so. Those beta-lactamase inhibitors on the market, such as clavulanic acid (which is combined with the antibiotic amoxicillin to form augmentin) only inhibit beta-lactamases that have a similar mechanism to serine proteases. No drugs yet exist to target the other class of beta-lactamases, which share their basic mechanism of action with metallo-proteases and which are the only ones that can break down the ‘beta-lactams of last resort,’ the carbapenems. ‘We are looking for compounds that inhibit a metallo-beta-lactamase known as NDM1, which has been found in Gram-negative bacteria that are resistant to almost every available antibiotic,’ says Jurgen Brem, a postdoc in Schofield’s group. ‘Working in the EU’s Innovative Medicines Initiative New Drugs for Bad Bugs programme, we have developed some extremely potent leads.’ The next step will be to test those compounds in animal models.
The university has spun out many successful companies, one of which, Oxford Drug Design, focuses on novel antibiotics. This company began life in 2001 as Inhibox to commercialise the work of chemist Graham Richards’ work in computer-aided drug design. It initially operated as a computational chemistry consultancy, and the change of name accompanied its switch to antibiotic discovery. ‘We have enhanced our basic cheminformatics technology specifically to investigate bacterial penetration, and are using it to design molecules that inhibit bacterial tRNA synthetase enzymes, and therefore prevent protein synthesis,’ says Paul Finn, the company’s chief executive.
Like bacterial ribosomes, bacterial tRNA synthases are sufficiently different from the equivalent mammalian enzymes to allow the design of selective inhibitors. They are not completely novel targets, but the only antibiotic targeting a tRNA synthase to have been marketed, mupirocin, is rapidly metabolised and can therefore only be used to treat skin infections. ‘Our best compounds inhibit the targeted synthetases with low nanomolar potency and kill the Eskape pathogens,’ adds Finn. ‘Studies in animal models of systemic infection have now started.’
Old drugs, new tricks
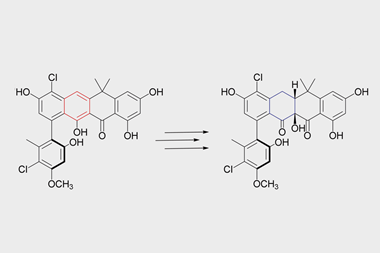
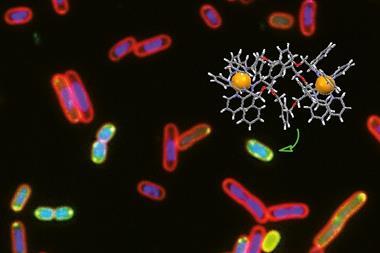
Drugs developed for one indication can sometimes be re-purposed for a different one. Between 2010 and 2013, a group of scientists at the University of Technology Sydney in Australia screened a large compound library including licensed drugs against drug-resistant Staphylococcus aureus and pulled out auranofin, an organogold compound that was FDA-licensed for the treatment of arthritis. This perhaps surprising discovery led to the spin-out of Auspherix, named to echo Australia as well as the chemical symbol for gold. The company is now based in Stevenage, UK, and is optimising a range of compounds based on a similar scaffold with a gold atom at the centre. These can kill many of the resistant bacteria on the priority lists, and, importantly, have been shown to disrupt bacterial biofilms. These consist of colonies of bacteria sticking together within a slimy extracellular matrix, often attached to a surface; they are resistant to most antibiotics and to the human immune response, and are therefore very difficult to eradicate. ‘We have defined a path from discovery into the clinic and will initially focus on compounds to treat complicated urinary tract infections,’ says Auspherix’s chief executive, Richard Rutter. ‘We already have a clinical development pathway planned for our molecules, but we still haven’t established the exact mechanism through which they kill bacteria.’
So far, CARB-X has awarded grants to companies from six countries representing three continents: the only Asian company in that exclusive list thus far is Bugworks, based in Bangalore, India. An announcement on investment in further Asian companies is expected soon. ‘We weren’t specifically looking for Indian companies, but for excellent science and development potential, and Bugworks ticked all the boxes,’ says Outterson. Scientists based there have developed a novel ‘stealth’ strategy for bypassing efflux pumps, preventing antibiotics from being ejected from bacterial cells. ‘Our strategy involves selecting effective molecules that are targeted by one particular efflux pump and designing out binding to the pump using structure-based methods, and we hope to take our molecules into Phase 1 trials in 2019,’ says Bugworks’ president of R&D V. Balasubramanian.
And there are many other companies with pre-clinical candidate antibiotics that seem at least as promising as those developed by Bugworks, Auspherix and Oxford Drug Design. These first stages of the pipeline may be fuller than they have been for years, but there are pitfalls ahead. ‘It has been estimated that only one out of 70 compounds in early development will ever be licensed,’ says Outterson. Small companies with public–private, ‘angel’ or charitable funding may take a molecule into early clinical trials, but later trials are more expensive and more challenging, and there is more at stake. Most people believe that those big pharma companies that have maintained antibiotic divisions, such as Merck, will have an essential role here, but they will still need to recoup their costs. What clinicians need most is ‘drugs of last resort’ to be used only in emergencies, but what company will invest millions in a drug with such limited sales prospects? It may be that a completely new funding mechanism is needed: perhaps an enormously generous one-off cash prize for developers that enables a drug to be distributed at cost price. Something like this already exists in the Longitude Prize for an accurate, affordable diagnostic test for bacterial infections.
So, are we any closer to outwitting resistance? To misquote Nobel-winning US molecular biologist Joshua Lederberg, ‘It’s up to us to use our wits to keep up with [bacterial] genes’. There is no longer such a shortage of novel molecules entering the antibiotic pipeline, but it will take all our imagination and political will – as well as a further step-change in funding, increasing it to the levels currently seen in cancer – to get enough of them through that pipeline and into the clinic. And all this will take many years. If we are to win the race against antibiotic resistance we will need to be even more careful with how we use the drugs we still have.
Clare Sansom is a science writer based in London, UK










2 readers' comments