Self-assembling superlattices built with DNA could usher in era of materials on demand
Two groups of scientists in the US have developed methods for constructing nanoparticle lattices from DNA bonds. Both techniques, based on either DNA origami or self-assembly, raise the prospect of chemically forging ‘materials by design’.
Since the 1980s, scientists have been manipulating DNA to create everything from walkers or rollers to exotic lattice patterns. Others have been exploring the idea that DNA will bind metal nanoparticles together in 3D lattices, preventing them from falling into the restrictive close-packed structures they typically adopt.
‘We are trying to create a whole new field of chemistry – we’ve been doing so for two decades now – where we define building blocks in a new way,’ says Chad Mirkin from Northwestern University, who won Chemistry World’s Entrepreneur of the Year award in 2013.
Back in 1996, Mirkin’s group realised single DNA strands could be grafted onto the surface of gold nanoparticles, which would bond together if they bore complementary strands.1 ‘You can think of a particle as an atom and the oligonucleotide as a highly programmable bond,’ explains Mirkin. ‘If things work well, you can begin to build structures that are highly ordered, in fact crystalline in nature…you can separate bonding characteristics from particle identity.’
In isolating identity from bonding, it appears the possibilities could be endless. ‘Once we know a particular structure can form, we can swap out any particle we like,’ says Mirkin. ‘As long as we use the same [DNA] sequences, it will assemble into the exact same structure.’
Bait and switch
But the team has now gone one step further, finding inspiration from a burgeoning field of biology to create a transmutable nanoparticle. ‘We were thinking of pluripotent cells,’ comments Mirkin. ‘The idea that…when introduced to appropriate chemical cues, they will differentiate into functional cells.’
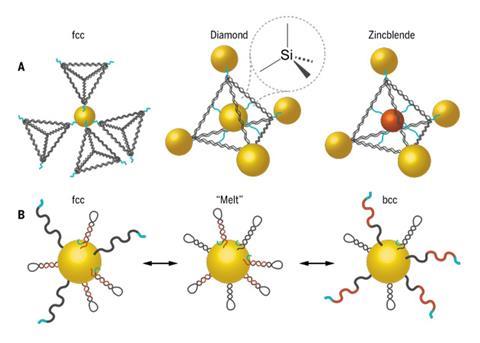
In their most recent study, the team’s gold nanoparticle, about 10–20nm in diameter, has a number of closed ‘hairpin’ DNA strands along the surface that cannot link with strands on neighbouring particles.2 ‘Effector’ strands, or short sequences of oligonucleotides, act as switches and open these hairpins, allowing the particles to bond into a lattice arrangement.
The transmutable aspect is apparent when different DNA strands are anchored to the particle. The switch can selectively open hairpins with complementary strands, allowing the particles to form a face-centred-cubic lattice. When non-complementary strands are exposed, a body-centred-cubic arrangement is favoured.
This crystal switching will allow the team to ‘program’ a whole host of crystal behaviours, from lattice parameters or crystal habit, and create structures that don’t even exist in nature. ‘There is no better example of crystal engineering on the planet,’ says Mirkin. ‘You’re effectively taking a particle without changing its core identity [and] you are changing its bond characteristics.’
Diamond league
Giving nanoparticles directional ‘valency’ is also being explored by Oleg Gang from Brookhaven National Laboratory and his colleagues.3 But their motivation stems from the pure challenge of creating complex structures like diamond from metal nanoparticles. ‘It turns out it’s an extremely challenging problem on its own from a structural point-of-view,’ says Gang. ‘In some sense the creation of diamond tests our ability to fabricate complex materials by self-assembly.’
Gang explains that the difficulty stems from the low volume fraction of diamond compared to other lattice arrangements. Structures will always tend to adopt a high density state.
Previous attempts to make diamond from DNA-coated metal nanoparticles have struggled with this issue and researchers often have to constrain the particle position and bonding parameters to a large degree. But these constraints aren’t necessary in Gang’s technique.
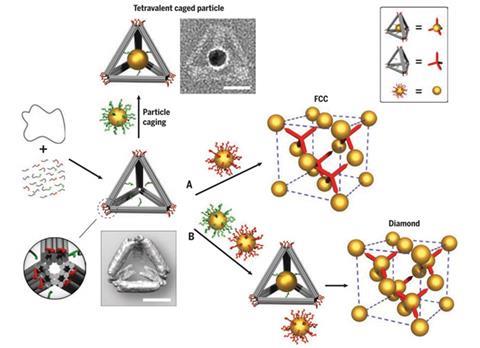
The team first made tetrahedral cages by folding an 8000-base long DNA strand containing ‘staples’ – short DNA strands that connect when brought together.
The cage has several ‘sticky’ single strands inside the open cage and bunches at each of the four outward-facing vertices. ‘Those strands that are inside, they essentially can capture the [DNA-coated] particles,’ explains Gang. ‘This particle is not isotropic any more, it can only interact according to those vertices.’ Caged particles can therefore coordinate with four neighbouring nanoparticles and the structure of diamond can be built.
Crystal control
For Gang, this self-assembly approach has a clear advantage over previous attempts: ‘You can establish the conditions where just mixing two particles, not really controlling how one will connect to another … [is] not constraining the system too much – you’re rather leaving a lot of freedom to the system and it can form diamond.’
This freedom is central to both Gang’s and Mirkin’s techniques and, in the latter case, Mirkin believes this freedom will open up new avenues in crystal engineering. ‘Nobody out there can, ahead of time when you make a new material, program crystal symmetry, lattice parameter and crystal habit – we can do that with this system,’ says Mirkin. ‘We’ve been trying to build, effectively, a new table of elements defined on these concepts that opens up a whole new field of materials-by-design.’
Jan Vermant, a soft materials chemist from the Swiss Federal Institute of Technology in Zurich, who was not involved in the work, is impressed by the research, but raises concerns about its scalability. ‘The control over structure is impressive,’ Vermant tells Chemistry World. ‘It will be interesting to see how these ideas and structures can be translated into materials with unique properties.’
References
1 C A Mirkin et al, Nature, 1996, 382, 607 (DOI: 10.1038/382607a0)
2 Y Kim et al, Science, 2016, DOI: 10.1126/science.aad2212
3 W Liu et al, Science, 2016, DOI: 10.1126/science.aad2080












No comments yet