Larger and more diverse DNA-encoded libraries on the cards thanks to new tool
Chemists in the US have designed a method that applies palladium-catalysed C(sp3)–H activation to DNA-tethered molecules. It provides a way to generate DNA-encoded libraries with staggering chemical complexity and diversity from relatively simple starting materials, creating new possibilities for drug discovery chemists.
Drug discovery programmes often focus on finding small molecules that can bind selectively to a specific target protein. This is a challenging task and involves tests on millions, or sometimes billions, of molecules in as few experiments as possible. Scientists can keep track of large numbers of molecules by attaching a unique DNA sequence to each drug candidate, with the specific order of DNA base pairs acting as a barcode to identify each individual molecule. The resulting collection of tagged compounds is called a DNA-encoded library. DNA-encoded libraries have expedited drug discovery programmes in recent years, enabling rapid identification of lead compounds. However, one of the key barriers DNA-encoded library users face is creating drug-like compounds in the presence of DNA.
Forming carbon–carbon bonds is a necessity for drug discovery chemists. In recent years, C–H activation has become an increasingly attractive approach for coupling carbon-based fragments. C–H activation typically uses a transition metal atom to cleave an inert C–H bond; it has created new ways of making organic molecules rich in sp3 nature, a favourable feature for drug candidates. ‘[The] C–H bond is a ubiquitous chemical bond in molecules and they are very strong. Breaking this bond opens new avenues to make molecules,’ says Jin-Quan Yu, from the Scripps Research Institute.
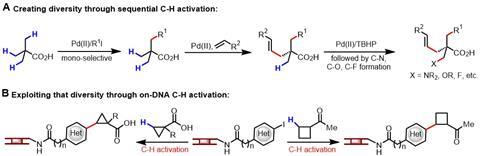
DNA-encoded library and C–H activation technologies have gained increasing attention amongst the scientific community in recent years, so it may seem surprising that no one has combined these techniques before. However, using C–H activation to functionalise a molecule that is bound to DNA presents three major challenges: Firstly, DNA solutions must contain water, a solvent typically incompatible with C–H activation protocols. Secondly, nucleotides within DNA are known to inhibit the transition metal catalysts used in C–H activation. And thirdly, the concentration of DNA-bound substrate is around 100-times lower than that typically used in an organic reaction. Yu’s group set out to merge the two in spite of these obstacles. ‘The combination of these technologies allows rapid creation and exploitation of molecular diversity in the context of drug discovery, though I must admit that this is also a fun challenge for us,’ says Yu.
Through careful optimisation of conditions that this group had previously developed for similar reactions, Yu’s team discovered a system that uses a palladium catalyst, a silver source and a base to perform C–H activation reactions on a wide range of DNA-bound aryl and heteroaryl iodides. The on-DNA substrates were coupled to carboxylic acids and ketone surrogates, which contained numerous pharmaceutically relevant functional groups. One of the key benefits of the methodology is that it introduces three-dimensional sp3 character into DNA-encoded libraries. Taking this a step further, the team used chiral amino acids as substrates, often attached to cyclopropane and cyclobutane rings, all of which are common features in successful drug compounds. And to fully exhibit the new methodology’s potential, the team also used it iteratively over three steps to create a complex drug-like compound from a DNA-bound heterocycle.
‘It’s quite exciting that we may now be able to call C(sp3)–H activation a process tolerated by DNA,’ comments Liam Hudson, an expert in DNA-encoded library technology at the Novartis Institutes for BioMedical Research, US. ‘The described work is fairly extensive with respect to the coupling partners, and suggests that novel libraries of good diversity and developability can be achieved through the application of creative design.’
Yu says the method could be expanded to encompass more types of molecule. ‘If the scope can be further improved, I think this will have major impact on drug discovery.’










No comments yet