
A bioluminescent nucleic acid sensor (Lunas) test has been developed that matches the speed and accessibility of a Covid-19 lateral flow test with the sensitivity of a PCR test.
Fast accessible disease diagnosis saves lives, as vividly illustrated during the Covid-19 pandemic. However even as tests for the disease became more accessible users were faced with a trade-off between PCR tests that require complex and expensive analysis that can take several days to a return a result, or a lateral flow test that yields results within minutes there and then but with potentially a thousandfold less sensitivity.
Maarten Merkx at Eindhoven University of Technology says that his team set out to address this compromise between speed and sensitivity for diagnostic tests. He explains that his group’s previous work was largely focused on fluorescent sensors, which have the drawback of needing light to excite the proteins into giving off light. This light can scatter making the target fluorescence signal harder to detect. In contrast bioluminescent proteins, like those found in fireflies, anglerfish and phytoplankton glow in the dark. People had thought the intensity of bioluminescence was generally too low for a high sensitivity device. However, this changed when Promega Corporation developed the bright stable bioluminescent protein NanoLuc.
The researchers used a split version of NanoLuc with one fragment attached to a type of Crispr-associated protein (dCas9) engineered to bind to a particular segment of the target DNA and the rest of the protein attached to a dCas9 protein that binds to an adjacent segment. Only when the two fragments bind to the target DNA does the whole NanoLuc protein assemble, allowing it to produce a detectable blue signal for picomolar (10-12 moles/litre) DNA concentrations.
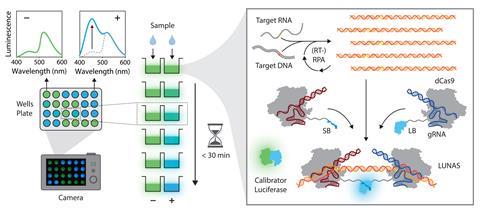
‘I was myself surprised that the signal was bright enough to be read by a smartphone camera,’ says Merkx. However, clinicians need even higher sensitivity, so Merkx and his colleagues introduced a recombinase polymerase amplification (RPA) reaction to increase the DNA concentrations without the complex steps required for PCR. With this the test yields a sensitivity in the femtomolar range (10-15 moles/litre).
However, the mechanism behind the protein’s bioluminescence posed an additional hurdle. As Merkx explains, the protein emits light by catalysing the oxidation of a substrate – an organic molecule added along with everything else. As the substrate is used up, the bioluminescence diminishes over time, making it hard to quantify the DNA load from the signal. Changes in other parameters such as pH or temperature can also affect the signal.
To get around this Merkx and colleagues added a second protein, consisting of NanoLuc tightly bound to green fluorescent protein. This green glowing protein was added whole, so it bioluminesces regardless of the presence of the target DNA but is still subject to the effects of substrate concentration, temperature and pH. Hence comparison of the blue and green emissions acts as a calibration device, a trick they pioneered with a previous bioluminescent protein marker sensor two years ago.
‘It’s a fantastic piece of work,’ says Nicolas Winssinger, who is deputy director of the Swiss National Centre for Competence in Research and specialises in organic chemistry and bioactive molecules. He points out that RPA is an exceptionally powerful amplification method but that only solves the detection threshold problem, not sequence specificity or readout. ‘The Lunas platform offers a practical and scalable solution to both problems,’ he adds.
Having already demonstrated it can match a PCR Covid-19 test’s sensitivity in minutes, they are now working on using it for other infectious diseases such as bird flu. They also plan to develop it further to detect multiple diseases with different coloured bioluminescent proteins.
References
HJ van der Veer et al, ACS Cent. Sci., 2023, DOI: 10.1021/acscentsci.2c01467
















No comments yet