A general-purpose, reprogrammable molecular computer has been constructed from DNA by a team of researchers in the US and Ireland. The system can execute different algorithms ranging from copying and sorting processes, generating random walks and executing cellular automata. It works by the self-assembly of DNA strands or ‘tiles’ into helices that form tubular structures by complementary base pairing. The emerging patterns on the tubes encode the output from the algorithm, and can be read out mechanically using an atomic force microscope (AFM) to inspect the molecular structures.
Computing with DNA relies on the fact that information can be encoded very concisely into the base pair sequence on strands. That alone has made DNA a focus of attention for very high density molecular memories, but the molecule can also be used to conduct computation. Early notions of DNA computation in the mid-1990s involved simple search processes in which all possible answers to a problem are encoded into DNA strands and then the correct answer is sifted out.1 But a more versatile approach, suggested in the late 1990s by Erik Winfree at the California Institute of Technology in Pasadena, and developed by several teams since the early 2000s, uses DNA self-assembly to embody an algorithmic process.2,3 It deploys DNA tiles that link via selective base pairing into arrays enacting a series of logic steps.
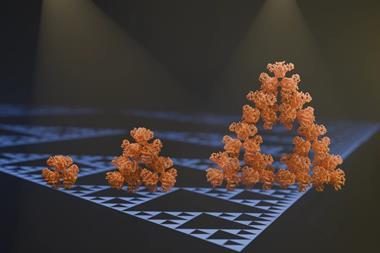
In this way, for example, Winfree and co-workers programmed DNA tiles to build themselves into a structure called a Sierpinski triangle, which repeats the same basic shape at ever larger scales in a fractal manner.3 Such computations involve tiles tailored to the task, however – very different from the general, reprogrammable tasks that regular silicon-based computers can conduct.
A bit of DNA
Now Winfree, in collaboration with computer scientists Damien Woods of Maynooth University in Ireland, David Doty of the University of California at Davis, and their colleagues, have made a set of DNA tiles that can be combined to carry out a wide range of computational algorithms – a big step towards a general-purpose molecular computer.4
At this stage the DNA computer juggles with only a handful of bits, doing nothing that would tax a regular computer. But as well as providing a proof of principle for making the system reprogrammable and scaling-up to a new level of complexity, already the results suggest how this kind of molecular computation might offer new possibilities for making materials and nanostructures.
‘It’s certainly a tour de force,’ says Nadrian Seeman of New York University, who pioneered DNA nanotechnology in the 1980s. ’They have taken algorithmic self-assembly to a new and higher level with what looks like high efficiency.’
The DNA tiles in this system embody the inputs and outputs of logic gates. They are single strands in which part of the sequence encodes two binary inputs – the four possible permutations 00, 01, 10 and 11 – and another part encodes two binary outputs. A circuit is built from these tiles via recognition sequences on each tile that determine how the tiles will clip together sequentially. Each two-input/two-output logic gate in the circuit is characterised by a ‘truth table’ that specifies the outputs for each pair of input bits, and the appropriate tiles are selected that will implement that table as they become attached to the growing array.
It’s a bit more complicated than that
In practice it is rather more complicated than that. For one thing, the system also needs other kinds of tile, which encode one-input gates. And each of these ‘abstract tiles’ is in fact represented by four real tiles, the redundancy building in a ‘proof-reading’ function that reduces errors in the computation due to occasional mismatching of tiles. These tiles assemble into helices that clip together at their edges into a cylindrical bundle of 16: a DNA nanotube that grows as the tiles assemble algorithmically in a physical embodiment of the computation. There is also a special tile that creates a ‘seam’ along the tube, which can be easily ‘unzipped’ using another strand that displaces this tile, turning the tube into a flat ribbon.
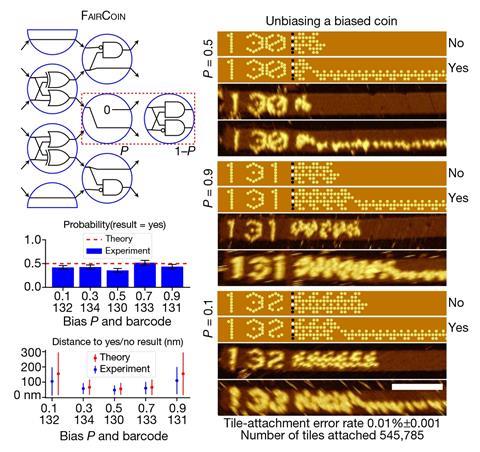
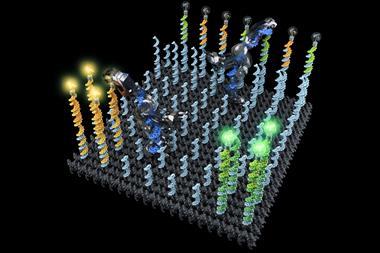
That process is crucial to the readout. The result of the computation, encoded in binary via the tile sequences, emerges after the self-assembly process has enacted the steps of the algorithm, and is then simply repeated in all subsequent growth of the tube. To reveal this binary result, the researchers attach the small organic molecule biotin to DNA encoding ‘1’ outputs, to which the protein streptavidin will bind in solution. This bulky molecule protrudes from the unzipped ribbons when deposited on a flat surface, so that it appears as a bright blob in a topographic scan with an AFM.
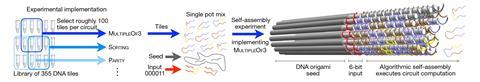
To carry out a computation the researchers select those tiles from the full set of 355 needed for the algorithm in question. They then seed a solution of these tiles with a nanotube nucleus marked with a ‘barcode’ – literally an alphanumeric number written into the initial DNA helical bundle using the biotin/streptavidin system that denotes the algorithm in question, easily visible in the microscopic readout. The tiles assemble over a day or two, the computation is completed, and the tubes are unzipped and the result read out. The researchers say that the error rate due to tile mismatching is less than 1 in 3000.
‘Very significant breakthrough’
The 355 tiles can encode all six-bit computations conceivable with this set – ‘more than enough to see lots of sophisticated computation’, says Doty. ‘The work provides a very significant breakthrough in DNA tile computation, allowing any algorithm using only six-bits to be compiled into a subset of their 355 tiles,’ says John Reif of Duke University in North Carolina, who has also worked on algorithmic computation using DNA tiles.
The researchers think that one early application could be in building complicated DNA nanostructures in a programmable way. ‘Our DNA tile set grew computationally sophisticated patterns, but it “painted” them onto a boring structure: a hollow tube that grows in one direction forever,’ says Doty. ‘Could we instead grow an object whose physical shape itself is controlled by the algorithm the tiles are executing?’ This, he says, would be a kind of ‘3D printing without the printer’.
‘I am fascinated by cases where the output of the algorithmic self-assembly is the shape of the structure itself,’ says Winfree. That would allow the construction of very complex molecular structures using a relatively small number of molecules, perhaps in a way that responds to the local environment. ‘Maybe some tiles could induce the tube to bifurcate into two branches, as in the development of a lung,’ Winfree says. ‘Different circuits or algorithms thus would correspond with different instructions to fabricate different structures.’
References
1 L Adleman, Science, 1994, 266, 1021 (DOI: 10.1126/science.7973651)
2 C Mao et al, Nature, 2000, 407, 493 (DOI: 10.1038/35035038)
3 P W K Rothemund, N Papadakis and E Winfree, PLoS Biol., 2004, 2, e424 (DOI: 10.1371/journal.pbio.0020424)
4 D Woods et al, Nature 2019, 567, 366 (DOI: 10.1038/s41586-019-1014-9)














No comments yet