Researchers build complex DNA structures that are larger than ever before
A collection of large, complex objects sculpted out of DNA have been unveiled by three separate research groups, expanding the range of nanometre-scale structures that DNA self-assembly can make. Groups led by California Institute of Technology’s (Caltech’s) Lulu Qian, Harvard’s Peng Yin, and Technical University of Munich’s (TU Munich’s) Hendrik Dietz have each developed complementary methods.
Shawn Douglas from University of California, San Francisco, who wasn’t involved in these studies, emphasises that the largest DNA structures now weigh billions of Daltons, and are a thousand times heavier than the largest were a decade ago. The tubes Dietz’s team constructs can also be up to 1000 nanometres long, ten times as big as the largest DNA structures were previously.
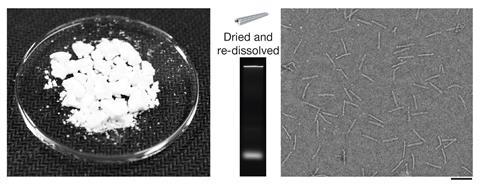
Dietz’s team has also devised a method for producing the DNA strands for €23 (£20) per milligram, less than a third of the current cost. And they say this figure could potentially be reduced to €0.18 per milligram. ‘In technology development, whenever you see a method’s power increasing by factors of 10, coupled with decreases in cost by factors of 10, it typically heralds some exciting applications on the horizon,’ Douglas says.
Remarkably, DNA construction is already at least 26 years old, from when New York University’s Ned Seeman published his group’s assembly of cubes from ten DNA strands in 1991. In the years since, researchers have built progressively bigger and more intricate DNA objects, and used them for computational and mechanical functions. One key underlying technology, known as DNA origami, relies on forcing one long scaffold strand of DNA into a desired shape using dozens of other, shorter, staple strands.
Dietz says, however, that the field has somewhat ‘stagnated’ in terms of applications. ‘Most…address niches in basic science and typically operate on the scale of micrograms or even less,’ he says. But he adds that learning to build larger, more complex structures may help address this.
Levelling up self-assembly
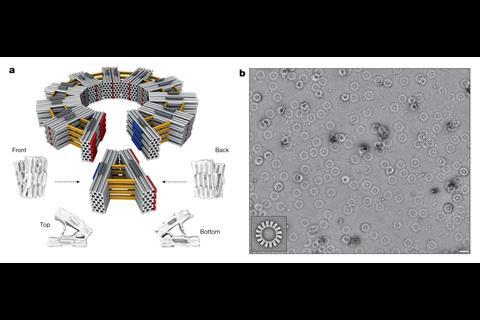
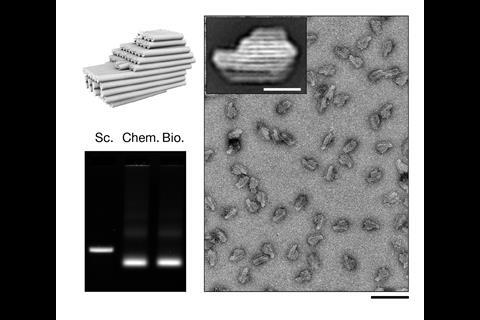
Dietz’s team was inspired by viruses, whose outer shells contain just a few types of protein subunit closed into regular shapes. They explored whether DNA origami subunits might do the same, trying different designs and studying their properties using cryo-electron microscopy. They discovered that these first-level subunits needed to form precise shapes and be rigid to successfully self-assemble at second and third levels.
‘The subunit needs to withstand collisions from solution molecules – the faces have certain relative angles and if they fluctuate too much they’ll never form a closed object,’ Dietz explains. They also shouldn’t bind too tightly, because they’ll get stuck in partially formed states. ‘If we have sufficiently weak interactions then subunits can associate but also dissociate. If you have some erroneously stuck subunits they fall off again.’
It is now conceivable to create much more sophisticated artificial molecular machines with sizes similar to that of a bacterium
Lulu Qian
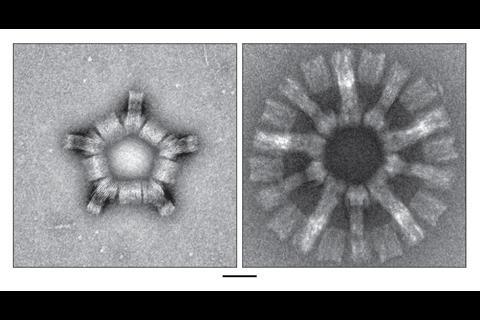
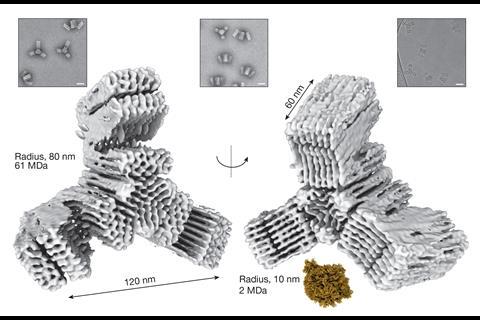
The TU Munich team’s final designs used V-shaped first-level DNA origami subunits, which could link up into second-level 350 nanometre diameter rings or ‘reactive vertices’.1 Depending on their shape, the reactive vertices could link up and close onto each other in third level virus-sized cages that were tetrahedral, hexahedral or dodecahedral. The largest weighed 1.2 billion Daltons and contained 220 DNA origami units. Similarly, the rings can link up into third level, 1000 nanometre-long tubes.
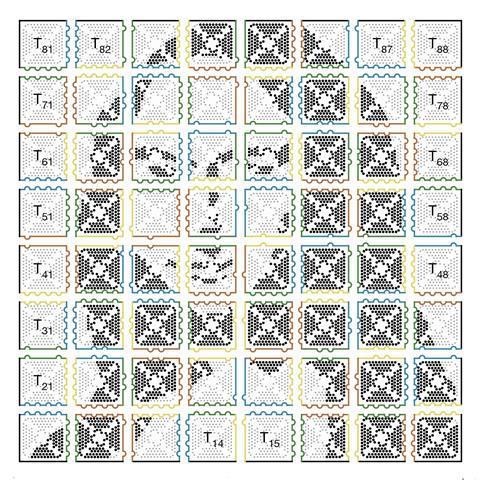
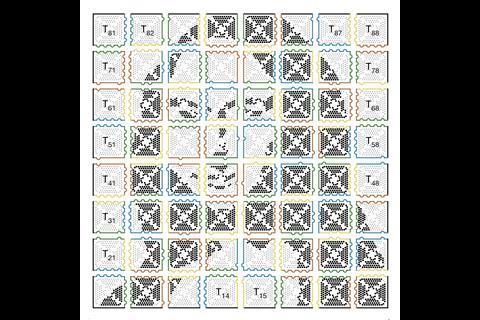
While TU Munich’s approach means all assemblies have to be symmetrical, the Caltech team’s multi-level assembly approach creates custom designs. They produce two-dimensional images from a jigsaw of 64 DNA origami tiles, reaching up to 8,704 pixels and 700 nanometres wide.2 ‘Once we have synthesised each individual tile, we place each one into its own test tube for a total of 64 tubes,’ explains Qian’s grad student Philip Petersen. ‘First, we combine the contents of certain tubes together to get 16 two-by-two squares. Then those are combined in a certain way to get four tubes each with a four-by-four square, and then the final four tubes are combined to create one large, eight-by-eight square composed of 64 tiles. We design the edges of each tile so that we know exactly how they will combine.’
Qian’s team calls this approach fractal assembly, comparing it to fractal objects where features repeat at different scales. Repeating design principles at different levels allows them to use the same small set of DNA strands for building structures of increasing size, with any possible pattern. ‘With fractal assembly, it is now conceivable to create much more sophisticated artificial molecular machines with sizes similar to that of a bacterium,’ Qian tells Chemistry World.
Pick and mix bricks
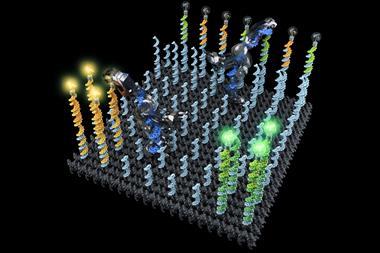
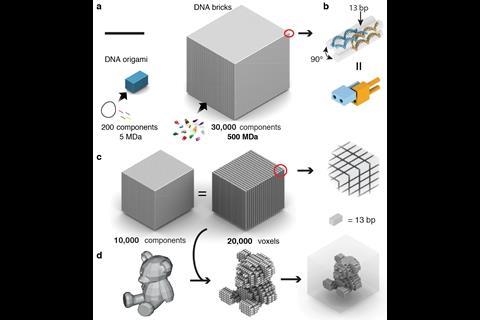
Unlike the Caltech and TU Munich teams, Peng Yin’s Harvard group doesn’t use DNA origami. Instead, his team builds on its 2012 ‘DNA brick’ work, which mixed 100 individual 32 base long strands in one pot to form solid cubes of DNA. Each strand is unique, with four different eight-base recognition areas, bending into U-shapes that interlock in specific positions among four other strands. ‘Each component only knows how to bind to four local neighbours,’ Yin explains. By omitting certain strands from these cubes, they leave gaps and form two-dimensional images and small three-dimensional objects.
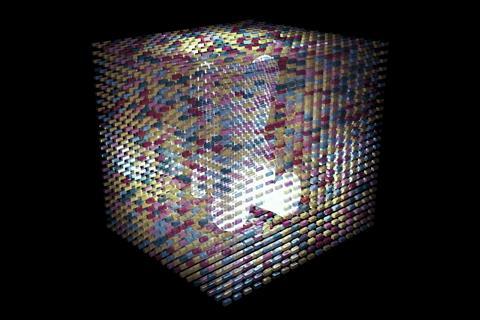
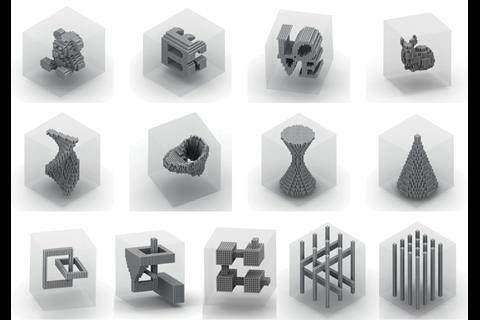
Yin’s team wanted to make larger three-dimensional shapes with cavities in the middle. After several years of work using longer, 52-base strands with four 13-base recognition areas, they were able to form such objects using 20,000 unique strands.3 Their final cubes measure 100 nanometres on each side, weighing up to a billion Daltons.
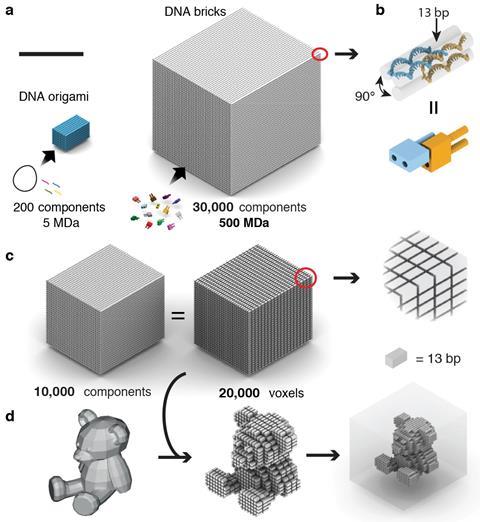
Yin imagines combining the fundamental principles in each of the groups’ approaches. ‘I’m hopeful we could make much larger and much more complex objects than what we demonstrate here, which already demonstrates a hundredfold increase in complexity,’ he says. ‘Maybe another tenfold and hundredfold capacity increase could be on the horizon.’
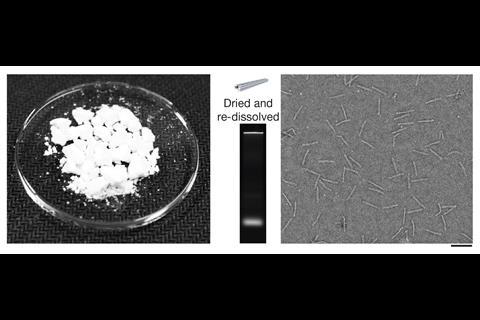
Dietz points out that for many labs, producing strands in such large quantities may be prohibitively expensive. To try and reduce the costs, his team recruited E. Coli bacteria to make hundreds of DNA staples along with DNA-based enzymes that can release them in one giant loop.4 They can do this by infecting the E. Coli with engineered bacteriophage viruses, which contain the blueprint for the loop. After growing the bacteria and extracting the DNA loops, adding zinc activates the enzymes and releases the staples.
These impressive milestones are only the beginning
Shawn Douglas
Although the team has only done it in a two litre lab-scale reactor, Dietz says the process should scale to thousands of litres, which should make exploring the capabilities of DNA nanostructures practical in new applications like drug delivery. Dietz stresses that there’s a lot of uncharted scientific territory because of the cost of producing large amounts of DNA. ‘You could embark into animal studies, to see whether we could put custom DNA objects to therapeutic uses,’ he says. ‘Quantity is really a big bottleneck in the field.’
Elisa Franco from University of California, Riverside calls the papers ‘impressive achievements in the field of DNA nanotechnology’. However, she would also like to see improvements to the folding speed because, although these papers don’t say how long it takes to form the final structures, it often takes several days.
Douglas agrees that there are more challenges to be solved, especially if such structures were to be integrated into applications like the DNA nanorobots he works on. ‘These impressive milestones may give the impression that the major challenges are solved, but they are only just beginning,’ he says. ‘I imagine the computing pioneers felt similarly when they finally had transistors working well enough for Gordon Moore to plot his famous extrapolation.’
Correction: On 8 December 2017 the units of one of the DNA structures was corrected
References
1 K F Wagenbauer et al, Nature, 2017, DOI: 10.1038/nature24651
2 G Tikhomirov et al, Nature, 2017, DOI: 10.1038/nature24655
3 L L Ong et al, Nature, 2017, DOI: 10.1038/nature24648
4 F Praetorius et al, Nature, 2017, DOI: 10.1038/nature24650




















No comments yet