Template-directed process that replicates non-biological molecules is step towards using evolution to explore chemical space
Scientists have developed a purely synthetic system for replicating information from one generation to the next. It copies information encoded in monomer building blocks from a parent, template sequence to a daughter sequence and is the first transfer of sequential information in a system that doesn’t interface with DNA.
In the search for materials with specific properties and functions, conventional methods use a time-consuming trial-and-error-based approach or optimise a known material. However, if materials were able to evolve like DNA, ‘you could find a way of selectively binding the molecule that has the best properties and then allow it to mutate, re-select the ones that bind and go through many iterations so that you evolve towards the property that you want,’ says Harry Anderson, an expert on template-directed synthesis from the University of Oxford, UK, who was not involved in the study.
‘In principle this is the answer to any problem that requires a molecule with a function,’ comments Christopher Hunter from the University of Cambridge, UK. Now, Hunter’s team has taken a step towards that goal with a general approach, powered by a covalent base-pairing system, for replicating sequence information in synthetic oligomers.
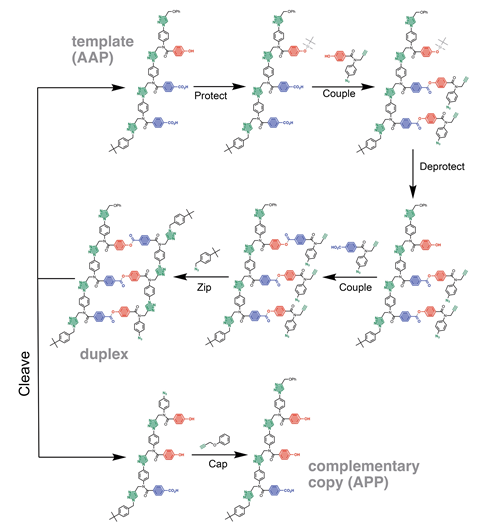
First, they generate the parent oligomer with a precise sequence of monomer building blocks. In this case, the monomers all have an alkyne and an azide group, and either a phenol or a benzoic acid unit. The azide and alkyne groups on the monomers allow a copper-catalysed cycloadditon reaction to zip them together to form the parent strand, with information encoded as the sequence of phenol and benzoic acid units. Now, the parent strand acts as a template for the complementary sequence. Phenol bases on the parent strand are selectively protected whilst ester base-pairing attaches phenol monomers to benzoic acid units on the parent strand. The protecting groups are removed before ester base-pairing attaches benzoic acid monomers to the phenol units on the parent strand, before another click reaction zips up the backbone. A hydrolysis reaction then cleaves the base-paring interactions to give the daughter strand.
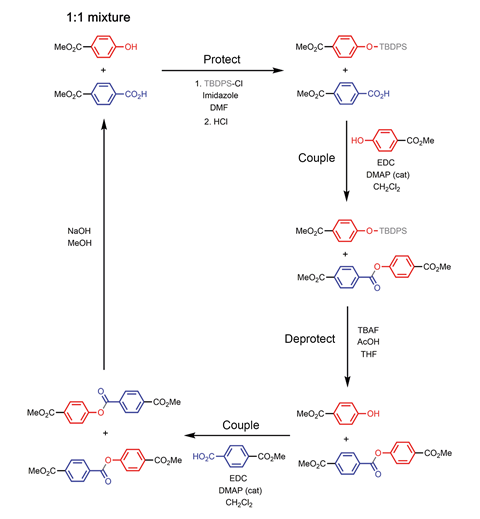
In previous research on template-directed synthesis, weaker interactions such as hydrogen bonding or reversible supramolecular processes like dynamic covalent chemistry bonded the monomer building blocks and template strand. ‘What is new and interesting about this approach is the use of covalent bonding to replace traditional DNA base-pairing,’ says Steven Zimmerman, who investigates supramolecular polymer chemistry at the University of Illinois, US.
The process does generate some side-products. ‘17% is a scrambled sequence. A bit of scrambling is precisely what you want for evolution as you would maintain the information that you have with a small amount of variation introduced,’ says Hunter. ‘The scrambled byproduct from out-of-sequence coupling illustrates the very high level of organisation required for efficient transfer of information,’ adds Anderson.
‘We’ve made no real attempts to optimise the ester base-pairing system,’ says Hunter. ‘The overall goal is to have a copying process that we don’t lose information in. It’s easy to see how you could make errors so getting something that is high fidelity in the first place is the real goal.’
References
This article is open access
D Núñez-Villanueva et al, Chem. Sci., 2019, DOI: 10.1039/c9sc01460h









No comments yet