The new online tool ChemMaps could help students and medicinal chemists explore chemical space in the search for new drugs. The site, which the developers dub ‘the Google Maps of drugs’, allows users to navigate a chemical universe of over 8000 drugs and 47,000 environmental problematic compounds in 3D. It enables users to search for a chemical and find out, based on its structure, what other types of similar compounds or drugs are on the market or undergoing testing in clinical trials.
‘I first thought about creating something like this a couple of years ago, when I saw that my colleagues, who are not computational chemists, were having a lot of trouble navigating, trying to browse, the chemical space,’ says Denis Fourches, an assistant chemistry professor at North Carolina State University who came up with ChemMaps.
‘I realised that in the real world, when people have to go somewhere they go to Google Maps and they easily browse it to go from one city to another city,’ Fourches says. ‘As a chemist, there was absolutely no tool available in the public domain to look at the structure of a chemical and see what kinds of chemicals or other drugs are very similar.’
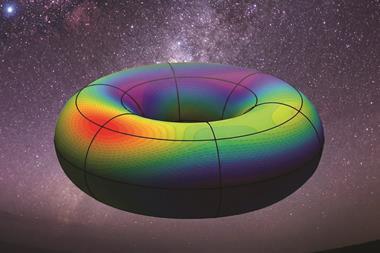
The compounds in ChemMaps appear like stars in the night sky – each molecule is represented in three dimensions and these coordinates are based on the properties of its chemical structure. When a user clicks on a particular compound, several key characteristics are displayed, including chemical structure, medical uses and other physical chemical properties. ‘A lot of properties are calculated for each chemical and when two points are close together it means that their structures are more similar,’ Fourches explains.

Because ChemMaps helps visualise the diversity in the neighbourhood of a particular compound or drug, it can also help with drug repurposing. A drug might be approved for a particular use, but the site helps reveal similar drugs that effectively treat other conditions, Fourches notes.
Chemical risk calculation
ChemMaps could also help improve chemical risk assessments. ‘When regulators receive compounds to be assessed for their toxicity, one of the many things they do is to look at the similarity of that compound to known toxicants,’ Fourches tells Chemistry World. ‘A tool like ChemMaps that easily projects your new compound into this map … can help the researcher understand why this compound is more or less likely to be toxic because of that chemical neighbourhood,’ he says.
Lee Cronin, who chairs the University of Glasgow’s chemistry department and works on constructing complex functional molecular architectures, is excited by ChemMaps. He says it provides ‘a nice visual tool’ to help chemists interact with large datasets and develop new ideas or research hypotheses.
Rafael Gomez-Bombarelli, a computational materials scientist at the Massachusetts Institute of Technology, agrees that ChemMaps can be quite helpful. ‘Chemical space is very complex and we need all the visualisations we can have to navigate it,’ he says. ‘We experience a three-dimensional world every day and this visualisation … can compress chemical space into the three dimensions we know and love.’
However, Gomez-Bombarelli points out that topographical maps only need to compress three dimensions onto two, but chemical space is ‘so high-dimensional and complex’ that there is no agreement about its equivalent for latitude, longitude and elevation. ‘Two chemical compounds can be very near each other, or very far away, depending on the choice of dimensions to express them,’ he says.
Derek Lowe, a medicinal chemist who blogs at In the pipeline, has similar concerns. ‘The idea of being able to click on a drug and see the closest neighbours in this artificial space is, in the end, of somewhat limited utility,’ he tells Chemistry World. Lowe suggests that the user would have to know more about the targets and binding sites of each drug, as well as its structure–activity relationship, in order to determine whether this connection is potentially interesting or just coincidental.
References
A Borrel, N C Kleinstreuer and D Fourches, Bioinformatics, 2018, DOI: 10.1093/bioinformatics/bty412
















No comments yet