5000 nanoscale experiments teach algorithm how to predict outcomes of reactions in the presence of inhibitors
The yields of tricky cross coupling reactions can now be accurately predicted by a computer program that taught itself how to tackle this tough problem. Key to the algorithm’s expertise is the data it trained on from thousands of small scale reactions. ‘The big goal, which this is a small step toward, is to be able to predict reaction performance of new substrates without experimentation,’ explains Abigail Doyle from Princeton University, who led the work together with Spencer Dreher from Merck & Co.
Machine learning has helped scientists explore chemical space, find new synthetic pathways and predict reaction outcomes. However, yield prediction software still often gets things wrong. This is because the data algorithms have to work with – reaction parameters collected by many groups over the years – is often inconsistent and incomplete. Reactions that don’t work, for example, are usually not reported.
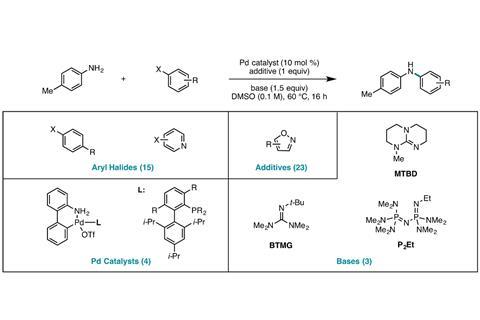
To overcome this issue, the US team created a bespoke database of almost 5000 Buchwald–Hartwig couplings, a palladium-catalysed reaction that makes a bond between carbon and nitrogen. An isoxazole – a heterocycle known to inhibit cross couplings – was added to each reaction. Despite this added difficulty, the algorithm the Princeton–Merck team trained on this data could correctly predict yields within a small margin, close to the experimental error.
As carrying out 5000 experiments would take a human chemist months or even years, Doyle and Dreher enlisted the help of Merck’s high throughput platform that can perform 1500 nanomolar reactions in a day. A random forest algorithm was then fed with the outcomes of 3000 of the reactions as well as calculated computational parameters – such as HOMO and LUMO energies – of each reagent.
The forest algorithm learns by building decision trees. ‘These trees could be yes/no questions like “Does the yield improve if the LUMO energy of the aryl halide goes up?”,’ Doyle explains. For each question, the program adds a new branch; the output is the average of thousands of decision trees.
To see how the algorithm’s prediction accuracy changes as it’s fed more data to learn from, the team tried training it on only 230 experiments. Although the model lost some of its predictive power, by one measure its accuracy changed relatively little. ‘I thought that the strong model performance with sparse data sets was particularly interesting since most groups would struggle to screen in excess of 4600 reactions,’ says Natalie Fey, a computational chemist at the University of Bristol, UK.
However, finding out the reasoning behind the algorithm’s predictions often remains challenging. ‘As the authors note themselves, the models can be very difficult to interpret,’ says Fey. Although this type of ‘black box’ method ‘may well be fine if the focus is on making a reaction work, but academically it is unsatisfactory’, she adds.
Nevertheless, ‘the outcomes [are] really promising’, says Anna Gambin, a computational molecular biologist from the University of Warsaw, Poland. ‘The great message from the paper is that, assuming the availability of the adequate predictors, classification of reaction efficiency is doable.’
The team hopes to teach their algorithm how to deal with more structurally complex compounds. ‘The substrates in our study are all flat,’ Doyle notes. ‘When you get to three-dimensional structures, that will pose additional challenges at being able to describe the differences between the substrates.’
References
D T Ahneman et al, Science, 2018, DOI: 10.1126/science.aar5169















No comments yet