Let’s watch how Crispr evolves into a platform technology
Thanks to the 2020 chemistry Nobel prize, Crispr–Cas systems will forever be associated with gene editing. But it’s worth mentioning that the system is also finding uses elsewhere, because at its heart is a DNA detector: an RNA guide that can be designed to hunt specific targets.
Coronavirus has become one of those targets. Identifying patients infected with Sars-CoV-2 is a bottleneck in many countries but it is central to strategies for containing and controlling the spread of the virus. Crispr-based nucleic acid detection methods have been explored within numerous diagnostic set-ups for a variety of infectious diseases, not just Sars-CoV-2. The field is still very much in its infancy but analytical tools that use Crispr already look set to be sensitive, specific, affordable, portable and easy to reconfigure for different genetic sequences.
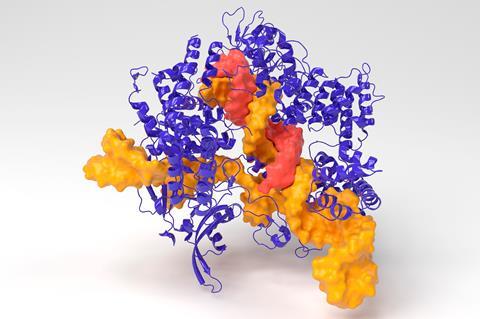
Another use of a Crispr–Cas system I’ve seen recently came from Douglas Clark’s team at the University of California, Berkeley in the US. Nature often organises enzymes in such a way as to channel reaction intermediates from one active site to another. These cascades boost reaction yields compared with analogous diffusional mixtures of molecules but mimicking how nature combines enzymes is tricky. Clark’s team has used a Crispr–Cas-based method to assemble a cascade containing five enzymes on a DNA strand by taking advance of how easy it is to customise what Cas binds to.1 They say the modular system is simple to programme and could work for virtually limitless types of enzymes.
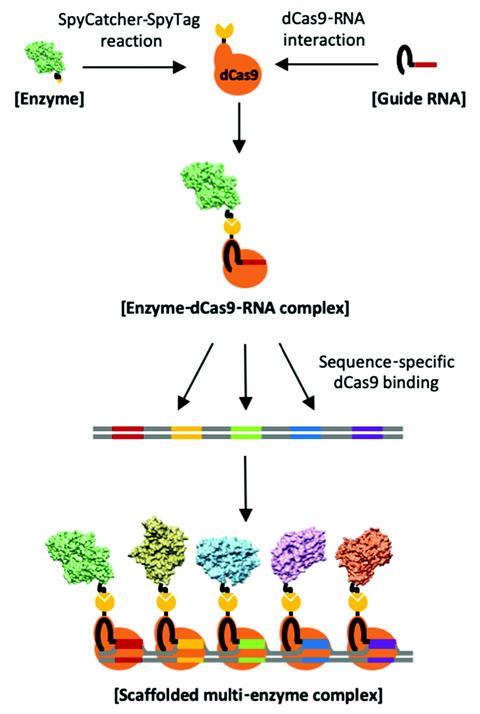
These are just two examples. Crispr was already topping the science barometer but I’m intrigued to see how it evolves as a platform technology now that is has a Nobel endorsement.
However, as the Crispr-based toolbox expands and becomes embedded within the scientific status quo there is a risk that originality and creativity get knocked down by the Crispr bandwagon. There may well be better gene-editing tools out there still waiting to be discovered, but now the motivation to find them is arguably much less. And I worry that labs rushing to adopt Crispr-based techniques encourages conformity rather than risk-taking.
Of course, it’s a testament to Crispr’s incredible potential that it carries such a price. But when Emmanuelle Charpentier and Jennifer Doudna teamed up to repurpose an ancient bacterial immune system they certainly weren’t following the herd. It’s important for humanity that scientific research maintains a delicate balance between exploiting a good idea and coming up with new ones.
References
1 S Lim et al, Chem. Commun., 2020, 56, 4950 (DOI: 10.1039/d0cc01174f)











No comments yet