Katrina Kramer tells the story of how Emmanuelle Charpentier and Jennifer Doudna developed the gene editing tool that won them the 2020 Nobel prize in chemistry
One of the greatest collaborations in biochemical history started in a café in San Juan, Puerto Rico. It was 2011, and Emmanuelle Charpentier and Jennifer Doudna were attending a microbiology conference that had a single small session dedicated to an obscure research field: clustered regularly interspaced short palindromic repeats, or Crispr for short. Just nine years later, the pair have shared the Nobel prize in chemistry.
Although the two scientists didn’t know each other, they decided to explore the city together after running into each other at a café. ‘We walked around the old streets of San Juan and talked about Crispr-Cas9 and how it would be really interesting to figure out what it might be doing in bacteria,’ Doudna recalled in a press conference last week. Discovered only a few years before they met, Crispr-Cas is part of an ancient bacterial immune system that detects and chops up invading viruses’ DNA.
Although Charpentier and Doudna had no plans other than to uncover the biochemical mechanism of this little-known system, they had a feeling that they were onto something big. Within a year of their accidental meeting, they would show how Crispr-Cas9 can be programmed to cut the DNA of any living cell with extreme precision – work that has revolutionised science.1
‘There’s no other tool that has been so revolutionary in the hands of scientists [and] in the lives of others, so quickly,’ says Helen O’Neill, who works on Crispr at University College London, UK. In the eight years since its inception, there has been an explosion of research activity that now sees tens of publications on the topic coming out every single day. Multiple companies have been founded solely to develop and commercialise the technology.
The genetic scissors have been used to produce blemish-resistant mushrooms with longer shelf life, mutant mosquitoes that could spread malaria resistance through wild populations and pigs whose organs could be transplanted into humans. Crispr-Cas could treat blood cancers and cure genetic diseases like congenital blindness or haemophilia.
There’s no other tool that has been so revolutionary in the hands of scientists and in the lives of others, so quickly
Unlike the decades-old concept of gene therapy, which tries to offset a mutation with an additional, healthy version of a gene, gene editing can entirely remove or replace a piece of disease-related DNA. In early clinical trials, Crispr-edited cells have eliminated the symptoms of another inherited disease, sickle cell anaemia. ‘The turnaround from it being a research tool to actually being a medicine was like a meteorite hitting the ground,’ says O’Neill.
And the technology continues to be used in innovative ways. Just a few weeks ago, a large team that included Doudna, and led by Melanie Ott at the Gladstone Institute of Virology and Immunology in San Francisco, US, showed that they could detect Covid-19 within five minutes using a Crispr-based test and a mobile phone.2
For developing the genetic scissors, Charpentier, from the Max Planck Unit for the Science of Pathogens in Germany, and Doudna, based at the University of California, Berkeley in the US, have now been awarded the 2020 Nobel prize in chemistry. But to understand how they managed to transform Crispr from a relatively obscure field of bacterial immunology to a worldwide phenomenon, we have to go back almost three decades.
From bacterial beginnings
In 1993, Francisco Mojica, then a PhD student at the University of Alicante in Spain, came across something unexpected in the genome of salt-loving microorganisms: repeating, palindromic sections of DNA separated by short spacer sequences. It was clear that these genes were active and important for the cells, but Mojica had no idea what they were for. Scouring the literature, he found a 1987 paper describing similar repeating DNA patterns in E. coli.3
After years spent digging through publications and examining prokaryotic genomes, Mojica realised that these strange repeating units were widespread in microbes – even among those that are far away from each other on the evolutionary tree. This indicated that the genes served an important purpose since they had been conserved across millions of years of natural selection. In 2002, Mojica, through correspondence with molecular microbiologist Ruud Jansen of Utrecht University in the Netherlands, coined the acronym Crispr: clustered regularly interspaced short palindromic repeats.4
But what did Crispr actually do? That remained a mystery until Mojica discovered that the spacers matched DNA sequences from phages, viruses that infect bacteria. The genes, he reasoned, must be part an undiscovered system that helps bacteria fight off infections.
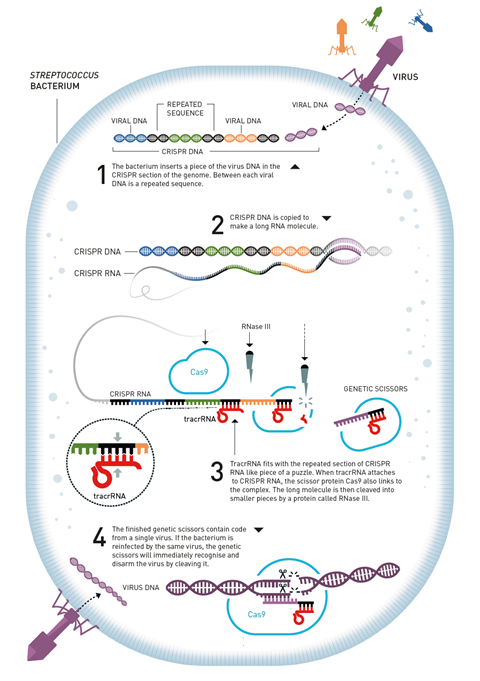
The scientific community was incredulous. Mojica’s study proposing a bacterial immune system was rejected by four different journals until he finally published it in 2005.5 In the years after Mojica’s publication, researchers slowly started piecing together exactly how this defence system works. The spacer regions are genetic clippings from phages, incorporated into a bacterium’s own genome after it survives a viral infection. Upon subsequent infection by a similar phage, the snippets are used to recognise and destroy viral DNA.
Crispr sequences are first transcribed into long RNA molecules, which are then chopped up along the repeating parts into small Crispr-RNAs (crRNAs), each containing a single spacer and a partial repeat. Several Crispr-associated (Cas) proteins are responsible for this activation process. The Cas proteins stay attached to the crRNAs, using them as a guide to find complementary viral DNA. Once a match has been found, Cas cuts the double-stranded phage DNA.
But there was still one piece missing from the puzzle. Many bacteria don’t have the multi-protein machinery that generates crRNA snippets, they only have a single protein – Cas9. There must be an alternative activation pathway, which Charpentier discovered in 2011.
Enter Charpentier
Charpentier came to Crispr research in the late 2000s. Having just moved from the University of Vienna in Austria to Umeå University in northern Sweden, she was studying Streptococcus pyogenes, the bacterium that causes tonsillitis and superficial skin infections but also pneumonia and life-threatening sepsis. Charpentier wanted to know out how this and other microbes develop antibiotic resistance by looking at small, gene-regulating RNA molecules.
Working with Berlin-based molecular microbiologist Jörg Vogel, Charpentier’s team had mapped all of S. pyogenes’ small RNAs. One variety stood out. It was clearly important, since it existed in large amounts, but nothing was known about it. The teams found its encoding genes sitting close to S. pyogenes’ Crispr sequence, suggesting some kind of connection.
It was a a beautiful story that was extremely unexpected and came right out of the blue
Experiments deleting different genes and seeing how this affected the microbe’s ability to defend itself against phages proved that these trans-encoded small RNAs – tracrRNAs – are an essential part of the process. TracrRNAs help generate the active, virus-destroying RNA–Cas complex. They bind to the repeat sections in the long RNA strand that is initially generated from Crispr genes. Cas9 recognises this RNA duplex and binds to it, triggering cleavage into smaller pieces by a ribonuclease enzyme to create the active crRNA–Cas9 system.
Charpentier, Vogel and their teams had found the missing activation pathway, which was based on an interaction between tracrRNA and crRNA. This was a radical idea – RNAs are not known to be team players, unlike proteins, which often work together.
After a year of collating evidence to support the hypothesis, Charpentier, Vogel and their teams published the results in 2011.6 Charpentier also presented them at a Dutch Crispr conference. ‘It was a highlight of the meeting – a beautiful story that was extremely unexpected and came right out of the blue,’ microbiologist and conference organiser John van der Oost said in a 2016 interview with Nature.
Charpentier knew she might be on to something big, but she needed a structural biologist’s help. So she attended the 2011 microbiology conference in Puerto Rico with a plan. ‘I had in mind to approach Jennifer and to ask her whether she would be interested in deciphering the structure of Cas9,’ Charpentier explained in an interview with The Crispr Journal last year.
From fun to eureka
‘When I first started working on it I thought of it as a fun little side project,’ Doudna said in a recent interview with the US National Institutes of Health (NIH) about her first foray into Crispr research back in 2006. Her Berkeley colleague, microbiologist Jillian Banfield, had told her about the mysterious Crispr sequences. Doudna was intrigued. But dabbling in Crispr research also made her feel a little guilty, she said. ‘I had funding from the NIH and the Howard Health Medical Institute, and I thought: How can I really justify this crazy little project on bacterial immunity that might have zero to do with human health?’
Just 10 years ago, Crispr was a relatively obscure field. The community was so small that everyone could fit into a single meeting room. ‘At that time, if I would go somewhere and say “I’m working on a bacterial immune system” I got very puzzled looks of “Why the heck would you do that?”’ Doudna recalled during a Berkeley press conference last week.
At the 2011 meeting in Puerto Rico, there was only a single session dedicated to Crispr. Both Charpentier and Doudna presented at this session. Although they knew of each other’s work – at that time, everyone in the field knew of everyone’s work – they had not met in person.
But as they started talking, they quickly realised their complementary scientific expertise. ‘We were coming to the Crispr field from quite different backgrounds,’ Doudna tells Chemistry World. ‘She is a microbiologist by training and focuses on bacteria that are harmful to humans. My research has always focused on molecules, and especially RNA and trying to understand its role in the evolution of life.’
In a 2017 interview with The Guardian, Doudna recalled Charpentier’s infectious excitement. ‘Even then I just had this gut feeling that this was something really interesting.’ As soon as the two were back in their respective labs – Doudna and her team on the US West coast, Charpentier and her group split between Sweden and Austria – they started working together.
‘There were lots of emails and Skype conversations and a lot of sending things back and forth across the Atlantic,’ recalls Martin Jinek, then a postdoctoral researcher in the Doudna group and joint first author on the seminal 2012 study. ‘Towards the end of the study, it essentially became an around the clock operation’, Jinek explains, due to the time difference between the three labs.
‘The collaboration was short and intense,’ Charpentier said in an interview with the Nobel Foundation soon after the prize announcement. ‘We understood that it was important to join forces and that we needed to go fast because the story was a great one.’
But at first, the experiments weren’t working. A combination of purified Cas9 protein and a piece of crRNA did precisely nothing. ‘We thought we had all the components, but the DNA was not getting cut,’ Jinek tells Chemistry World. ‘Then, as soon as we added the tracrRNA into the mix, suddenly things started to work. This was, for us, one of the big moments.’ The tracrRNA, it turned out, didn’t just play a role in activating the system, it was also needed for its smooth running.
‘By definition, Cas9 is a programmable enzyme: it’s using different sequences of RNA to recognise DNA,’ explains Doudna. ‘We then realised that we could probably design a single guide RNA that would work for programming Cas9.’
And that’s exactly what they did. They fused crRNA and tracrRNA into a single molecule – a 20 base pair sequence now called guide RNA. When it worked just as efficiently as the individual molecules, the teams realised that they had created a system that would cut DNA wherever they wanted by simply changing the RNA template – a eureka moment. ‘Martin was in my office and we were talking about his data, and we looked at each other and realised that this could be an extraordinary tool in other kinds of cells because of its ability to trigger DNA repair and thereby enable genome editing,’ Doudna recalled during the Berkeley press conference.
The idea was revolutionary – maybe too revolutionary for some. At roughly the same time that Charpentier, Doudna and colleagues worked on publishing their results, a team led by Virginijus Šikšnys at Vilnius University in Lithuania also showed that the Cas9 system could cut DNA sequences at a predetermined place. Šikšnys sent the paper to Cell, where it was rejected without review. It took several months until it was finally accepted at a different journal.8 Charpentier and Doudna had more luck, and their seminal study was accepted in Science.1
‘We presented the results close to the time of publication at the Crispr conference, and the excitement was huge,’ says Krzysztof Chylinski, the other joint first author on the paper, who was a PhD student in Charpentier’s group in Vienna at the time. While genetic engineering wasn’t new, Crispr-Cas would allow scientists to manipulate DNA with a whole new level of precision while making the process simpler, quicker and cheaper.
From years to months
‘People started thinking about genome engineering from the time when they realised what DNA is and that there is a code for life,’ says Audrone Lapinaite who recently completed a postdoctoral fellowship in Doudna’s lab and is now running a research team at Arizona State University in the US. When Crispr was published, there were two major players in the genome editing field: zinc finger nucleases and transcription activator-like effector nucleases (Talens).
‘The biggest problem with those two systems was that it was not that easy to target them to specific genomic sequences, because you had to re-engineer proteins that find the genomic location of interest and perform the cutting,’ explains Lapinaite. ‘That engineering takes a lot of effort, and it’s not trivial. With Crispr, the big breakthrough was that the protein is always the same, you can easily produce a lot of it.’
Before Crispr, says O’Neill, ‘it took me two years – and that was a great achievement – to get two transgenic mice. Now that could be done in a matter of months.’ Scientists now only need to produce a custom RNA molecule – a routine procedure – that matches the DNA sequence they want to target.
The only restriction on the freedom to cut DNA anywhere is the presence of a protospacer adjacent motif (Pam), a short sequence of two to six base pairs near the target sequence. The Pam is important for Cas9 to recognise its target. It stops bacteria from cutting up their own DNA as the Pam they use are specific to phages. But since Cas proteins from different bacterial species recognise different Pam sequences, even this restriction is slowly disappearing.
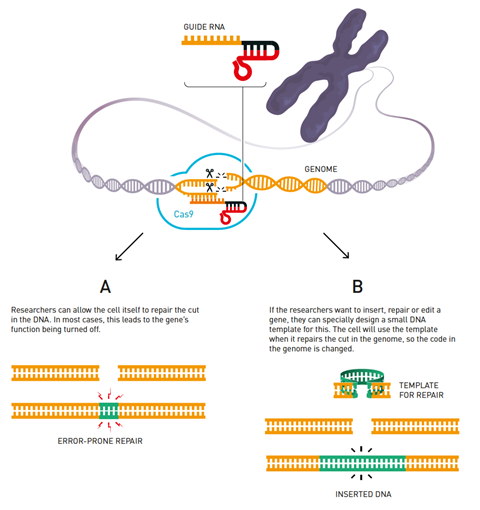
Crispr-Cas9 system can only cut double stranded DNA, but not alter it. The actual editing happens afterwards, relying on the cell’s own repair mechanism. Cells can repair DNA by directly fusing the broken ends back together, which usually deletes or inserts a few base pairs and tends to switch off a gene’s function.
In a different process, the cell looks for a homologous DNA sequence to use as a template for repair. Researchers can exploit this mechanism to insert or alter the sequence at the break by offering the cell small DNA template pieces. Since repair mechanisms are only present in dividing cells, editing the genes of non-dividing cells, neurons for example, remains challenging.
The idea spreads
‘What was for me a big sign of what was going to come was that, just within half a year, Feng Zhang and George Church showed that what we proposed is working indeed,’ says Chylinski. At first, it wasn’t clear whether Crispr-Cas9 would work in non-bacterial cells. One of the initial ideas was to simply use the system to make bacteria more phage-resistant, which in itself would have been a huge boon for biotechnology. But Zhang, Church and Doudna herself quickly showed that there were no such limits to Crispr-Cas9, demonstrating that the genetic scissors worked just as well in mouse and human cells.9,10,11
Over the next few years, the field experienced a flurry of activity that shows no signs of abating. ‘We didn’t quite appreciate how much hunger there was in the community for tools that could be readily applicable,’ says Jinek. Some scientists started modifying crop plants to make them more resistant to disease and pests. Others were engineering mice and rats to act as models for human diseases, while many jumped on the possibility to address genetic disorders.
But being able to edit genomes with such ease and precision also meant that it became possible to make changes to organisms’ germline, meaning cells that pass their genetic material on to their offspring. This was first demonstrated12 in monkeys in 2014 and then a year later in non-viable human embryos.13
‘Changes in the human germline, if taken far enough, would really change the course of human evolution,’ Doudna explained in the NIH interview. ‘I found myself lying awake many nights thinking about the potential for this technology that I’d been involved in developing being utilised in this fashion,’ she said in a 2019 talk about the future of gene editing. Ever since 2014, she has been publicly advocating for the responsible use of gene editing.
The ethical conundrum
But in November 2018, Doudna received an email with the subject line ‘Babies born’. It came from He Jiankui, a biophysicist at the Southern University of Science and Technology (SUSTech) in Shenzhen, China. ‘He told me in a very terse email that he had been involved in a clinical study where they used Crispr-Cas9 to make changes to the genome of babies who had actually been born,’ she recalled in the 2019 talk.
It was very, very important that the international community come out strongly against the reckless use of Crispr
A few days later, Jiankui publicly announced that he had disrupted the CCR5 gene, which allows HIV to infect cells, in twin girls. This was apparently in an attempt to protect the babies from the virus as they were born to an HIV-positive father. But while natural CCR5 mutations do provide some HIV protection, the gene edits were different and had never even been tested in animals.
‘I was pretty shocked,’ Doudna tells Chemistry World. ‘I knew that this was something that would attract a lot of attention and that it was very, very important that the international community come out strongly against that reckless use of technology that’s not responsible to people’s health.’ Indeed, the scientific community roundly condemned Jiankui’s actions and in December 2019 he was sentenced to jail for three years for illegal medical practice.
The scandal put gene editing into the spotlight. Public approval cooled to the point that more people were worried about the technology than excited about its possibilities. This has also affected the research community which was looking to help prevent people with devastating genetic illnesses passing them on to their children. ‘It’s set people like me, who had genuine honest intentions for that therapy, back so far, it was actually quite a frightening time,’ recalls O’Neill.
At the same time, ‘there’s been a really good concerted global effort to ensure that we don’t see that kind of reckless use again’, says Doudna. Not only national but worldwide discussions are needed, says Lapinaite. Last year, the World Health Organization launched a registry to track human genome editing activity.
Progress despite patent disputes
More positively, Crispr research has seen its first clinical success stories. Last year, Victoria Gray became the first person with a genetic condition – sickle cell disease – to be treated with a Crispr-based therapy in the US. She received a bone marrow transplant containing around two billion of her own cells, edited to switch back on a foetal haemoglobin-producing gene to compensate for her defective adult haemoglobin gene. Although it is too soon to fully evaluate the treatment’s success, Gray has stopped experiencing bouts of excruciating pain, which are typical for the condition, and has required no more blood transfusions.
There is an enormous drive to develop Crispr for such clinical applications but also as a biomedical tool. Both Charpentier and Doudna have each founded several startup companies focused on commercialising the technology.
But the rush to bring Crispr-Cas to market has also sparked a rather complicated patent dispute between on one side the two laureates and Berkeley, and Zhang and the Broad Institute on the other. Although Berkeley filed for a patent earlier – for using Crispr-Cas9 in cell-free systems – the Broad’s patent for using the technique in eukaryotic cells was granted first. So far, the two patents have been allowed to remain standing side by side, but the dispute is ongoing.
Meanwhile, the field is pushing ahead on many fronts. One of the challenges is delivering Crispr-Cas into specific cells. Traditionally, the entire system was introduced as an encoding piece of DNA, which enables the cell to produce its own Cas9 proteins and guide RNA. However, this floods the cell with a huge number of Cas9 proteins, increasing the chances of off-target activity. Scientists have started to test whether virus-like particles can inject the fully assembled guide RNA–Cas9 complex into cells. This allows better control over the amount of Cas protein per cell, and could even target specific cells types depending on the virus particles’ receptors.
Researchers are also exploring Crispr-Cas variations,14 some with different activities for RNA-cutting or single-strand nicking, for example. The latest peak of precision are DNA base editors, biochemically modified Cas enzymes that convert a single base pair into another without cutting the DNA strands. This could help treat genetic conditions caused by a point mutation such as progeria.
And people keep discovering new Crispr-Cas systems. Recently, the Doudna group described a tiny CasΦ they found in huge phages.15 ‘These bacterial viruses tend to have Crispr systems of their own, to kill off other viruses that are competing for the same host,’ explains Basem Al-Shayeb, joint first author on the study. Although only half the size of Cas9, the Crispr-CasΦ system is just as functional. And being that small could make it easier to deliver into cells, adds Al-Shayeb.
From collaboration to celebration
After their 2012 collaboration, Charpentier’s and Doudna’s interests diverged. ‘Emmanuelle was keen to continue really focusing on pathways in bacteria that are responsible for disease in humans,’ explains Doudna. ‘And in my own work, we dug into the molecules and mechanisms of Crispr.’
Nevertheless, the two remained in the centre of the Crispr craze and were showered with high-profile awards, including the $3 million (£2.3 million) Breakthrough prize in life sciences in 2015, the $1 million 2018 Kavli prize in nanoscience (which the pair shared with Šikšnys) and the $100,000 2020 Wolf prize in medicine. In 2015, Time magazine named them among the 100 most influential people.
Charpentier and Doudna were also regularly tipped for a Nobel prize and led prediction polls for several years. ‘As much as it was said by scientists – and non-scientists – that Crispr-Cas would be awarded the Nobel prize one day, it wasn’t clear to us,’ Charpentier said in a press conference following the Nobel announcement. ‘I was very emotional when I received the call.’
While Charpentier received her call from Stockholm sitting in her office in Berlin, Doudna’s was a less traditional wake-up call. It was 2.53am in California and her phone was buzzing. ‘It was Heidi Ledford, a Nature reporter I’ve talked to in the past,’ Doudna told the Berkeley media team last week. Ledford wanted a comment on the Nobel prize. ‘I thought she was calling to ask me to comment on someone else winning the prize. So I asked her: “Who won it?” And she said: “Well, you!”’
Charpentier and Doudna stress that Crispr is a perfect example how solutions come from unexpected places. ‘It’s not about doing science for the sake of publishing and accumulating impact factors, it’s about believing in a project that may not be as exciting or fashionable at first,’ Charpentier said in the press conference.
The 2020 chemistry prize was also the first to go to an all-female team. ‘It means a lot to me, thinking back to my own childhood,’ Doudna said to the Berkeley team. ‘Growing up, I have been told, more than a few times, that girls don’t do chemistry.’ In the Nobel press conference, she added: ‘I think for many women there’s a feeling that no matter what they do, their work will never be recognised as it might be if they were a man. I’d like to see that change and I think this is a step in the right direction.’
But what Doudna is most excited about is the dedicated parking space that Berkeley reserves for Nobel laureates. ‘Now I can finally, after 18 years, park on campus,’ she laughed.
References
1 M Jinek et al, Science, 2012, 337, 816 (DOI: 10.1126/science.1225829)
2 P Fozouni et al, MedRxiv, 2020, DOI: 10.1101/2020.09.28.20201947
3 Y Ishino et al, J. Bacteriol., 1987, 169, 5429 (DOI: 10.1128/jb.169.12.5429-5433.1987)
4 R Jansen et al, Mol. Microbiol., 2002, 43, 1565 (DOI: 10.1046/j.1365-2958.2002.02839.x)
5 F J M Mojica et al, J. Mol. Evol., 2005, 60,174 (DOI: 10.1007/s00239-004-0046-3)
6 C Pourcel, G Salvignol and G Vergnaud, Microbiology, 2005, 151, 653 (DOI: 10.1099/mic.0.27437-0)
7 E Deltcheva et al, Nature, 2011, 471, 602 (DOI: 10.1038/nature09886)
8 G Gasiunas et al, Proc. Natl. Acad. Sci. USA, 2012, 109, E2579 (DOI: 10.1073/pnas.1208507109)
9 L Cong et al, Science, 2013, 339, 819 (DOI: 10.1126/science.1231143)
10 P Mali et al, Science, 2013, 339, 823 (DOI: 10.1126/science.1232033)
11 M Jinek et al, eLife, 2013, 2, e00471 (DOI: 10.7554/eLife.00471)
12 Y Niu et al, Cell, 2014, 156, 836 (DOI: 10.1016/j.cell.2014.01.027)
13 P Liang et al, Protein Cell, 2015, 6, 363 (DOI: 10.1007/s13238-015-0153-5)
14 K S Makarova et al, Nat. Rev. Microbiol., 2020, 18, 67 (DOI: 10.1038/s41579-019-0299-x)
15 P Pausch et al, Science, 2020, 369, 333 (DOI: 10.1126/science.abb1400)



















1 Reader's comment