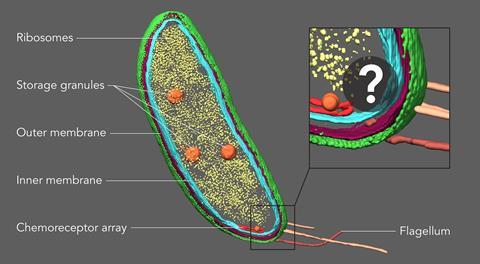
Inside the crowded confines of a cell, individual proteins can now be pinpointed with exceptional accuracy thanks to the combination of two powerful microscopy techniques.
Advances in cryogenic electron tomography (CET) produce images at near atomic scale resolution using purified proteins. However, in order to view a protein in its natural environment, a cell, CET is less useful. ‘One of the real challenges, is you’re not looking at a purified protein complex anymore,’ explains Peter Dahlberg, a postdoctoral researcher at Stanford University.
Within a cell there are a multitude of proteins that interact with each other and other cellular structures like membranes and ribosomes. When viewed using CET it can be difficult to tease out what’s what within the cytoplasm, meaning most proteins of interest for biologists aren’t directly discernable. This is where Nobel-prize winning super-resolution fluorescence microscopy comes in, as it is a powerful tool to pinpoint proteins.
Combining the methods allows a protein to be viewed within a cell. For this, samples are imaged fluorescently first and then with CET, after which the images are combined. However, this poses a problem.
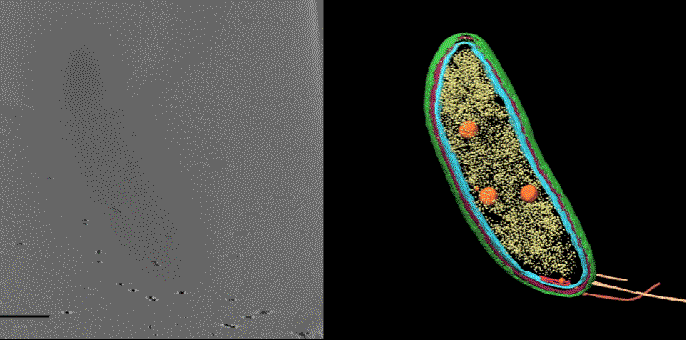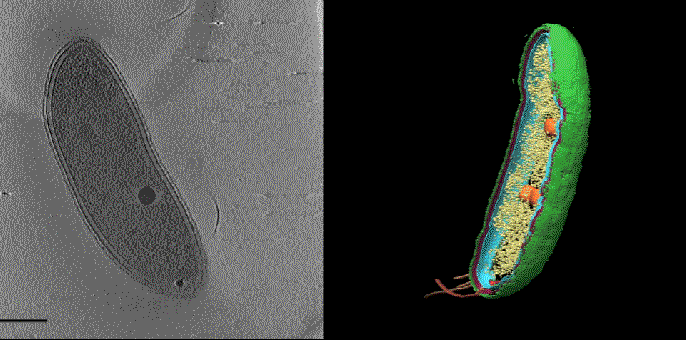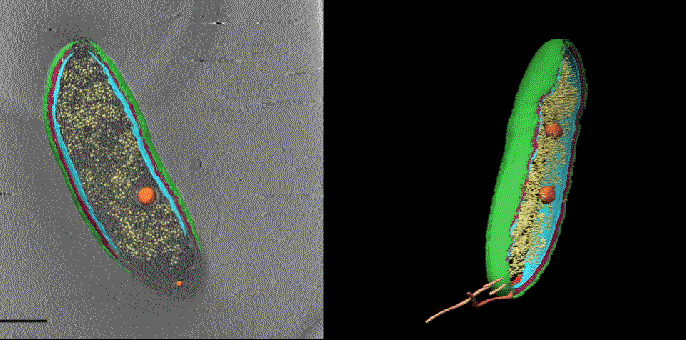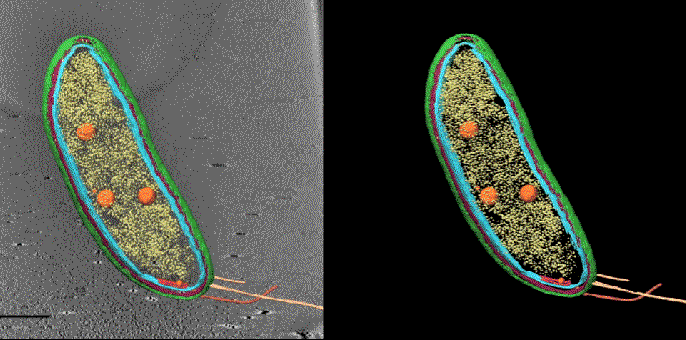
To prevent shifting between the two sets of images the samples are frozen at cryogenic temperatures. Getting fluorescence images with the same levels of resolution as CET is normally done at room temperature and doesn’t work as well in frozen samples. To improve the performance of super-resolution fluorescence at low temperatures Dahlberg leveraged his lab groups 2018 discovery of a red photo activated protein called PAmKate that maintains its fluorescence at cryogenic temperatures. ‘PAm was a big step forward for us in being able to do this, but it’s also a lot of brute force and making something that’s not ideal work,’ he explains. Experiments with the new technique, dubbed correlated imaging by annotation with single molecules (CIASM), showed that it could achieve localisation of proteins within the bacteria Caulobacter crescentus, on average, to 10nm.
Tâm Mignot, a bacterial cell biologist at Aix-Marseille University, says this work is a nice proof of concept that addresses a long-standing problem. ‘You see where the protein is by fluorescence, but now you can really look at its localisation related to the membrane or sides of the membrane, which you could not achieve in the past.’
Providing localised context could prove useful for the study of diseases caused by bacteria, viruses and parasites, all of which use unique proteins and structures to invade and exploit host cells and tissue. Identifying these improves our understanding of diseases and might provide targets for drugs that disable their vital processes. Malaria, for example, creates protrusions on the surface of red blood cells which cause them to get stuck in blood vessels. According to Friedrich Frischnknecht, a parasitologist at the University of Heidelberg, electron tomography of these protrusions reveals a spiral structure, ‘but nobody knows what the spiral is made from’. Using a technique like CIASM might provide an answer as to how this spiral structure, which has a direct link to pathogenicity, is built.
While the technique provides a more precise location of where a protein is, extracting and directly visualising that protein is not yet possible. Another challenge, adds Dahlberg, will be increasing the numbers of proteins that can be localised using this technique to the same levels found using room temperature methods.
References
P D Dahlberg et al, Proc. Natl. Acad. Sci. USA, 2020, DOI: 10.1073/pnas.2001849117









No comments yet