A machine learning platform has been used to discover new antibacterial compounds. The system screened more than 107 million chemical structures and found nine potential antibiotics, with one in particular showing potency against 97% of the drug-resistant bacteria it was tested against.
Aiming to tackle the growing antibiotic resistance crisis, a collaboration of synthetic biologists and computer scientists developed a deep learning platform to predict antibiotic activity. In particular, the group wanted to discover compounds with structures distinct from known antibiotics, to boost their chances of success against drug-resistant bacteria.
‘We wanted to demonstrate that these techniques are sufficiently strong to be used to identify therapeutic molecules,’ says Massachusetts Institute of Technology (MIT) computer scientist, Regina Barzilay, whose team developed the machine learning tool. Barzilay and her colleagues teamed up with researchers from synthetic biologist James Collins’ lab, who hoped to see if AI could help address the threat of antibiotic resistance.
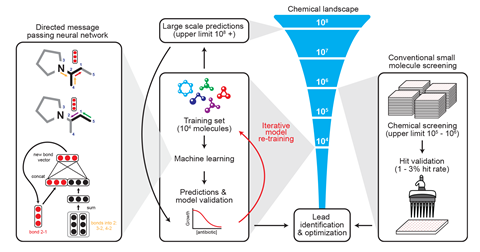
Collins explains that the machine learning model was trained using a library of 2500 molecules, comprising 1700 Food and Drug Administration approved drugs and another 800 natural compounds, which had been tested for activity against Escherichia coli.
‘The model that we developed learns to translate molecules into a continuous vector. It starts by having every atom represented with a vector of simple properties like the type of atom and whether it is in a ring and so on,’ says Barzilay. This is used to build up a fingerprint of the molecule’s structure, which coupled with the results of the E. coli tests, helps the neural network ‘learn’ molecular features that are associated with antibacterial activity.
Antibiotic hit
The model was then applied to the Broad Institute’s drug repurposing hub – an open access library of more than 6000 molecules with known biological activity. ‘It contains compounds at various positions in the drug development pipeline,’ explains Jonathan Stokes, who worked on the project at MIT. ‘So, it contains more or less the human pharmacopoeia of drugs, as well as molecules in late preclinical, and then phase one, phase two and phase three clinical trials.’ From this they discovered a compound, halicin, which has impressive antibiotic activity, despite having a chemical structure unlike conventional antibiotics.
‘Halicin turns out to be a remarkably potent novel antibiotic, with broad activity against a range of antibiotic-resistant pathogens,’ says Collins. ‘We applied it to 36 different panels of multidrug-resistant bacterial pathogens from the [US Centers for Disease Control]. And halicin was effective against 35 of those 36.’
The team then tested halicin in two different mouse models. ‘The first mouse model was an Acinetobacter baumannii skin infection model,’ explains Stokes. A. baumannii is a key target for medical researchers, as it acts as a reservoir for antibiotic-resistance genes and survives for prolonged periods on surfaces. ‘So, we establish an infection on the back of mice and then topically treat it with halicin,’ says Stokes. ‘And in 24 hours, we were able to eradicate the infection.’
The second mouse model demonstrated that halicin could clear a Clostridium difficile gut infection within four days.
Needle in a haystack
Following this success, the team applied their AI technique to a database known as ZINC15. The database contains more than 1.5 billion compounds and of these, 107 million were selected for screening based on physical properties known to affect biological uptake.
Based on the deep learning tool’s predictions, 23 compounds were chosen for further investigation, with eight displaying antibacterial activity against a range of pathogens. Two of these compounds showed promise against a range of drug-resistant E. coli.
‘It is great to see the application of machine learning to the search for new antibiotics whilst taking into account [adsorption, distribution, metabolism, excretion],’ says University of Birmingham microbiologist Laura Piddock, who directs scientific affairs at the Global Antibiotic Research and Development Partnership. Piddock is impressed with the proof of principal data for halicin, and highlights its low predicted toxicity in humans.
Katie Simmons, an expert in virtual high-throughput screening based at the University of Leeds, UK, describes the halicin results as ‘stunning’. ‘What is even more amazing is the ability of deep learning to select antibacterial compounds from such a large subset of the ZINC15 database,’ she adds.
‘It is unfortunate that research into new antibacterials has largely fallen out of favour with “big pharma” but the approaches described here could go some way to helping address our need for new therapies,’ says Simmons.
While the predictive power of the deep learning tool is impressive, Barzilay notes that it still requires scientists’ knowledge and experience. ‘These methods are becoming stronger and stronger, but they’re still not at the point when we can say that in silico screening can fully substitute experimental high-throughput screening,’ notes Barzilay. ‘Even if you have an imperfect predictor, it’s still useful for identifying interesting patterns and correlations that you can take advantage of. But for that you need the knowledge of medicinal chemists and biologists,’ she says.
Barzilay adds that further work is underway to improve the accuracy of the deep learning tool’s predictions. Her team are also developing AI methods that can be used beyond an initial screen – so that after identifying a lead molecule, the model will then suggest modifications that could improve its biological activity. Meanwhile, Collins’ lab aims to complete preclinical work on halicin and then potentially partner with industry to get it into clinical trials.
References
J M Stokes et al, Cell, 2020, DOI: 10.1016/j.cell.2020.01.02














No comments yet