Semi-synthetic organism shows there is nothing unique about the chemistry of life as we know it
The capacity of life to use alternative biochemistries has been demonstrated by researchers in the US, who have shown for the first time that the workhorse bacterium Escherichia coli can be engineered to use non-natural base pairs in its DNA to make proteins containing non-natural amino acids.1
While the replication of DNA containing non-natural base pairs has been reported before, this is the first time that the increased information they make available has been used in living cells.
It raises the possibility that life on other worlds might not use the same biochemical basis as ours.
‘It has long been argued that the molecules of life are somehow special and different from other types of molecules,’ says Floyd Romesberg of the Scripps Research Institute at La Jolla in the US. ‘We’ve demonstrated that this may not be true. Our man-made part, the unnatural base pair, can function in what is arguably the most central of all biological processes: the storage and retrieval of information.’
The work is also a step towards creating drugs and organisms with totally new functions. ‘The immediate goal that really drives us is to use the semi-synthetic organism to create new classes of protein drugs,’ says Romesberg. ‘Protein drugs have revolutionised medicine, but the properties that they have, and thus the diseases that they can be developed to treat, must in some way be limited by what they are made of.’ He hopes that his approach can broaden that repertoire.
Translation task
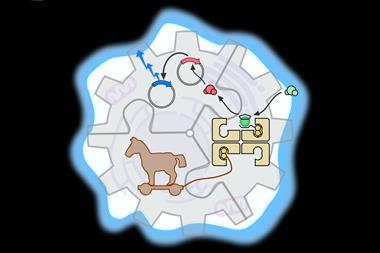
Earlier work by Romesberg and other groups showed that non-natural base pairs can be inserted into the DNA of living organisms, but using them to write and read out genetic information is another matter. Genes are translated into proteins via messenger RNA (mRNA) and transfer RNA (tRNA), which come together at the multi-protein enzyme called the ribosome to link up amino acids held by tRNA into proteins.
Each amino acid is carried by a different tRNA, the code being read out by matching of a three-nucleotide codon on the mRNA with a complementary anticodon on the tRNA. So to read out the information in a non-natural pair of nucleotides, the complementary partners must be incorporated into both the mRNA and tRNA and function at the ribosome. This is what Romesberg and colleagues have achieved.
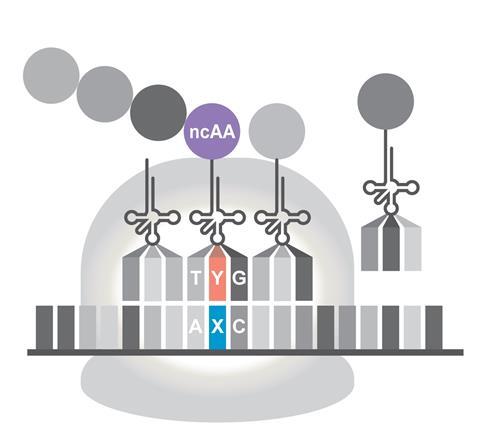
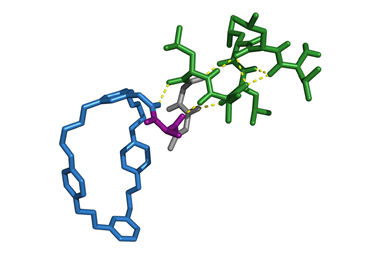
What is particularly remarkable about their alternative nucleic-acid chemistry is that their new base pair isn’t linked together, like normal DNA bases, by hydrogen bonding. Instead, it incorporates bulky organic heterocyclic groups which stick to one another in water by the mutual attraction of hydrophobic units. The Scripps group has showed that a bacterial tRNA engineered to have an anticodon with one component of the non-natural pair can recognise and stick to the complementary codon in mRNA transcribed from a corresponding gene within a DNA plasmid added to the cells.
A crucial trick is to use tRNA-loading enzymes that are not sensitive to their non-natural anticodons, such as the E. coli tRNA synthetase that loads tRNA with the amino acid serine. The researchers first used this system to insert serine at a position on a standard protein – green fluorescent protein (GFP) – coded by the non-natural base pair.
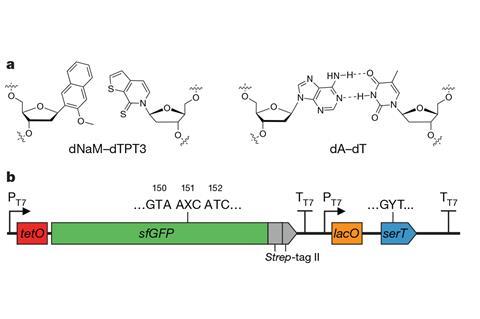
Then they got more ambitious, using the same approach to insert non-natural amino acids into the protein. First they replaced the gene for the serine-tRNA synthetase with one that makes an enzyme used in a different microorganism, Methanosarcina barkeri, which uses a rare amino acid pyrrolysine. The researchers used this tRNA synthetase to add a non-natural pyrrolysine derivative to GFP. As a further illustration of the expanded amino acid alphabet, they also used a tRNA synthetase from the archean Methanococcus jannaschii to make GFP containing the markedly unorthodox amino acid p -azido-phenylalanine.
The immediate goal that really drives us is to use the semi-synthetic organism to create new classes of protein drugs
Floyd Romesberg, Scripps Research Institute
This system gives a big boost to the amount of information that can be genetically encoded. The unnatural base pair makes available 152 new codons, says Romesberg’s Scripps colleague Yorke Zhang. ‘As we continue to develop it, it could be used to simultaneously and efficiently direct the incorporation of as many unnatural amino acids as you like,’ he says. The team is now working to demonstrate the simultaneous insertion of four different unnatural amino acids into a protein.
‘This is an important step in technology development toward expression of proteins with non-natural amino acids,’ comments Eric Kool of Stanford University in the US, who has also worked on expanding the genetic code. ‘If the technology ultimately enables the high-efficiency expression of proteins with multiple unnatural amino acids, it could be very important for future protein therapeutics and biotechnologies.’
References
Y Zhang et al, Nature, 2017, DOI: 10.1038/nature24659










No comments yet