Gene transplant repositions algal hydrogenase so that it directly captures electrons from photosystem I
Scientists have successfully directed photosynthetic electron flow away from fixing carbon dioxide and towards proton reduction by fusing together photosystem I (PSI) and algal hydrogenase in vivo. Modified algae cells expressing the PSI–hydrogenase chimera produce hydrogen in a light dependent fashion at high rates.
While hydrogen is a promising alternative to fossil fuels, currently 95% produced in the US is from steam methane reforming, which still generates greenhouse gases. Powered by the sun and using water as the electron source, producing hydrogen in algae has the potential to be a sustainable alternative but does come with its own challenges.
Although algae can naturally produce hydrogen, the production rate is much too low to be commercially viable. Previous attempts to improve the process have been hampered by oxygen inactivating the hydrogenase and other processes within the cell competing for electrons.
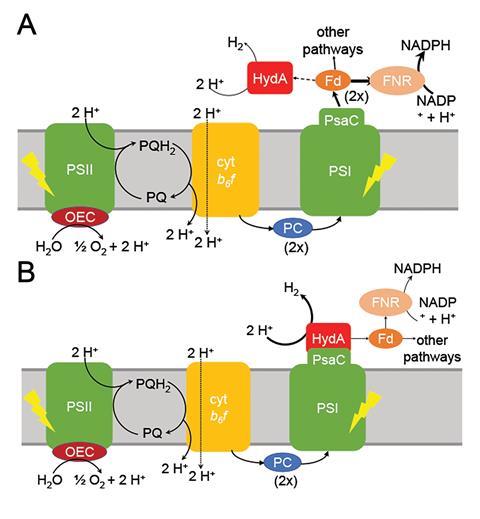
However, the PSI–hydrogenase chimera, developed by Iftach Yacoby from Tel Aviv University in Israel and Kevin Redding from Arizona State University in the US, and their colleagues, overcomes the electron competition problem by repositioning algae’s natural hydrogenase enzyme to directly capture electrons from photosystem I before they enter the general cellular pool. ‘We figured out a way that you could intercept these electrons coming from photosynthesis, at the place where they’re the highest energy so that you can drive interesting chemistry,’ says Redding.
The scientists took the entire structural gene for the hydrogenase (hydA2) of green algae Chlamydomonas reinhardtii, usually found in its nuclear genome, and transplanted it into the sequence for the PsaC subunit of photosystem I. They selected the turn of a β-hairpin in PsaC as the optimal insertion point for the hydrogenase, corresponding to residues 32–36 of PsaC. The psaC–hydA2 gene was then introduced into the chloroplast genome to replace the endogenous psaC gene. This results in modified algae cells capable of producing a dihydrogen every ~6ms in vivo.
Jenny Zhang, an expert in semi-artificial photosynthesis at the University of Cambridge, UK, says that identifying the site on photosystem I where redox enzymes can be fused is an exciting step forward. ‘This will no doubt form the foundation of future efforts to develop algal systems that can efficiently perform a range of solar-driven chemical formation processes, such as the conversion of carbon dioxide into useful feedstocks. The development of such breakthrough systems are still greatly needed in our repertoire of energy production and carbon dioxide recycling strategies.’
‘We are now moving towards trying to find hydrogenase enzymes that are more oxygen-resistant,’ says Redding. ‘The hope is that by using directed evolution, we could force the algae cells to do all this in the presence of oxygen, so that in order to live, they would have to make hydrogen.’
References
This article is free to access until 22 June 2020
A Kanygin et al, Energy Environ. Sci., 2020, DOI: 10.1039/c9ee03859k










No comments yet