DropSynth makes hundreds of strands in a single tube, boosting tests of gene function
Biotechnologists have been frustrated by the cost of making large pools of genes – but now US researchers can cheaply make collections of thousands of different DNA strands. Sriram Kosuri’s team at the University of California, Los Angeles (UCLA), calls its approach DropSynth, and say it produces each gene for less than $2 (£1.48). Existing commercial suppliers charge at least $25 for the shortest real genes the team has produced. ‘An individual in a lab can make 10,000 genes,’ Kosuri tells Chemistry World. ‘Before this paper was published that would have taken $1 million and a consortium to build.’
The UCLA team is amongst the researchers creating the demand for large DNA pools, using such collections in ‘multiplexed reporter assays’ over the past five years. These assays insert many genes into different cells that have been modified so effects on gene function can be read by high-speed next-generation, or next-gen, sequencing.
We could easily now synthesise all the genes for a particular antibiotic target and see which ones are resistant
Calin Plesa, UCLA
A single scientist can therefore explore thousands of small changes in gene sequences and their effect on biological functions such as protein production. For example, UCLA team member Calin Plesa now wants to use DropSynth to feed into assays that will study antibiotic resistance. ‘We could easily now synthesise all the genes for a particular antibiotic target, test them and see which ones are resistant,’ he says.
Kosuri has previously worked on microarray-based synthesis techniques that have already reduced costs for producing individual strands to a few cents per DNA letter. He assembled genes that are hundreds of letters long from smaller oligonucleotide chunks, known as ‘oligos’. The oligos have sequences of letters at each end that recognise the other chunks they’re supposed to combine with.
When mixed together these recognition sequences pair up in the same way that DNA does in its iconic double helix shape. Kosuri and his colleagues originally made copies of the final genes using the well-established polymerase chain reaction (PCR) to amplify them to get enough for their experiments. ‘That was great, but the problem was that if you build 10,000 genes you still have to do 10,000 amplifications in parallel,’ Kosuri explains, which is time consuming.
Milking it
Now, Plesa and his colleague Angus Sidore have found a way to assemble hundreds of oligos and amplify them in a single tube, emulating next-gen sequencing. To read many genes in a single tube, next-gen sequencing attaches the genes to beads and surrounds the watery mixture they’re in with oil. By rapidly spinning these mixtures, geneticists can then form milk-like oil and water emulsions, isolating the beads in the watery droplets to enable sequencing without interference from other genes. Plesa and Sidore similarly load beads to each carry all the oligos needed to make a single gene (see video). The UCLA scientists can therefore use an enzyme to release the oligos into isolated milky droplets, and then assemble and amplify them through PCR without interference.
The UCLA team developed the approach to where it could produce a few hundred genes in high purity and initially wrote a paper on that basis. However, an anonymous scientist reviewing their manuscript questioned whether it could really be scaled to produce thousands of genes. ‘We said, “OK, we’re going to make 10,000 genes”,’ Kosuri recalls, ranging from 381 to 669 letters long. ‘It took two weeks.’
To demonstrate feeding their pools into multiplexed reporter assays, they also cultured 1152 of the genes they made in E. coli whose native genes had been knocked out. ‘Using these assays with the DropSynth library as an input and sequencing as a readout, you can get a very interesting understanding of how each gene functions,’ Plesa says. ‘That’s becoming increasingly important because we have all the sequence information, we just don’t know what most of it is doing,’ Kosuri adds.
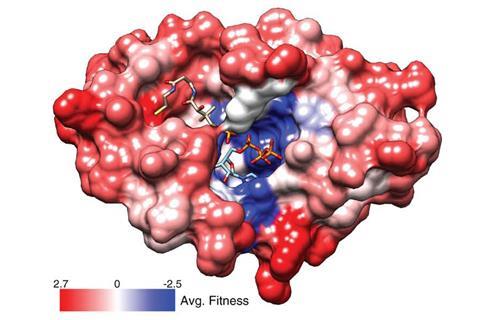
‘DropSynth complements existing high-throughput sequencing and genome editing technologies,’ comments Andrew Ellington from the University of Texas, Austin. He adds that it should now help scientists to create artificial sequences and assess their ‘fitness’.
Ali Tavassoli from the University of Southampton, UK says that ’the ability to pull down the three to five oligos required for a given version of a gene from a pool in parallel with those required for other variants, and assemble and amplify each of these into a functional gene is impressive.’ He adds: ‘The complexity of the technique may limit its adoption by academic labs that currently use error-prone PCR for the preparation of mutagenesis libraries. But current methods cannot be employed for the parallel preparation of gene libraries, which is where the real potential for this method lies.’
















No comments yet