The science behind the 2017 Nobel prize in chemistry
The 2017 Nobel prize in chemistry has been awarded to three scientists ‘for developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution’.
The laureates are: Jacques Dubochet from the University of Lausanne, Switzerland; Joachim Frank from Columbia University, New York, US; and Richard Henderson from the Medical Research Council Laboratory of Molecular Biology in Cambridge, UK. They each contributed to developing a technique that allows scientists to see the intricate structures of proteins, nucleic acids and other biomolecules, and even study how they move and change as they perform their functions.
What is cryo-EM?
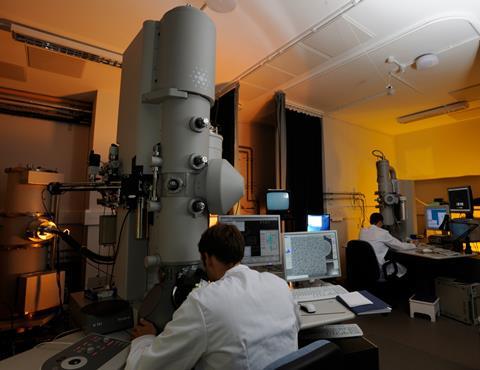
Transmission electron microscopes (TEMs) use a beam of electrons to examine the structures of molecules and materials at the atomic scale. As the beam passes through a very thin sample, it interacts with the molecules, which projects an image of the sample onto the detector (often a charge-couple device; CCD). Because the wavelength of electrons is much shorter than that of light, it can reveal much finer detail than even super-resolution light microscopy (which was awarded the chemistry Nobel prize in 2014).
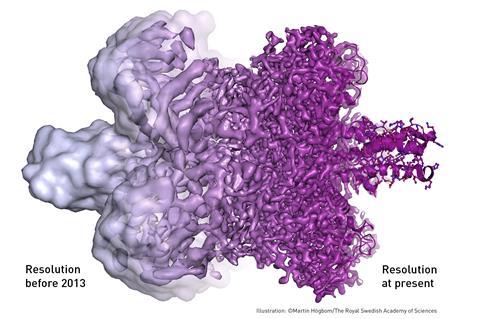
But some materials – particularly biomolecules – are not compatible with the high-vacuum conditions and intense electron beams used in traditional TEMs. The water that surrounds the molecules evaporates, and the high energy electrons burn and destroy the molecules. Cryo-EM uses frozen samples, gentler electron beams and sophisticated image processing to overcome these problems.
But can’t you look at proteins using x-ray diffraction or NMR?
X-ray diffraction can give very high resolution structures of biomolecules, and several Nobel prizes have been awarded for just that. But to get an x-ray structure, you need to be able to crystallise the molecule. Many proteins won’t crystallise at all. And in some cases, the process of crystallisation alters the structure, so it’s not representative of what the molecule looks like in real life.
Cryo-EM doesn’t require crystals, and it also enables scientists to see how biomolecules move and interact as they perform their functions, which is much more difficult using crystallography.
NMR can also give very detailed structures, and investigate conformational changes or binding of small molecules. But it’s limited to relatively small proteins or parts of proteins, and usually soluble intracellular proteins, rather than those embedded in cell membranes. If you want to study larger proteins, membrane-bound receptors, or complexes of several biomolecules together, cryo-EM is where it’s at.
What did the laureates contribute?
Each of this year’s three winners solved part of the problem of looking at biomolecules in a TEM.
Richard Henderson was the first person to generate an image of a protein using TEM. He packed many copies of the purple light-harvesting protein bacteriorhodopsin into a sample and combined images from multiple angles, using a low-power electron beam, to generate a 3D image of the protein. He continued to refine these techniques over several decades until he could produce images at the same resolution as those from x-ray diffraction.
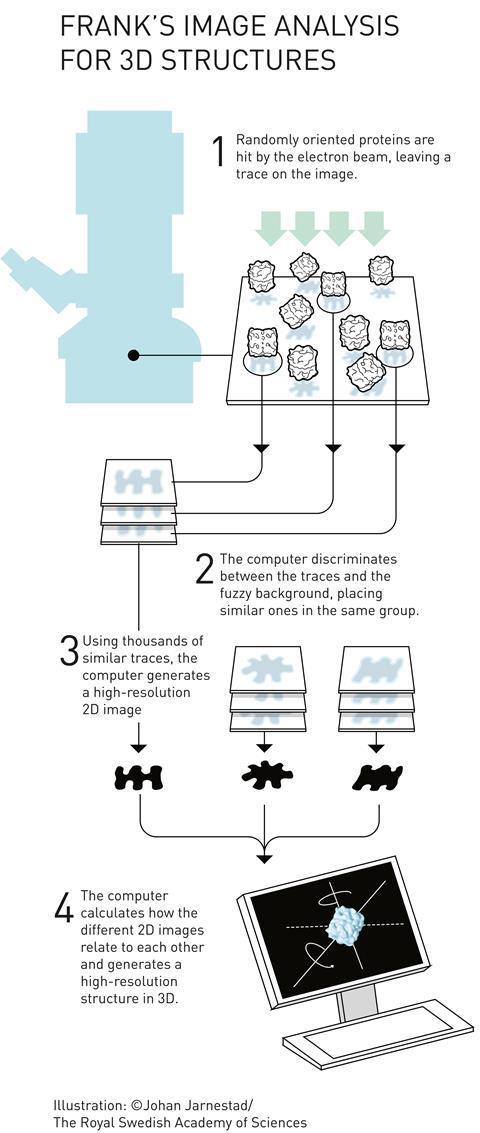
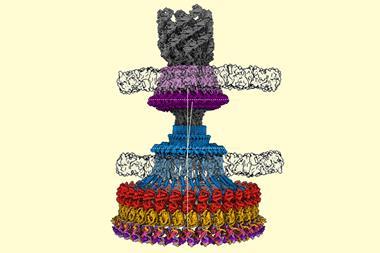
Joachim Frank’s major contribution to the field was in processing and analysing cryo-EM images. He developed computational methods for taking images of multiple, randomly-oriented proteins within a sample and compiling them into sets of similar orientations to obtain sharper 2D images. He could then construct a 3D image from these 2D projections. He used his algorithms to generate images of the ribosome – a massive structure made of several proteins and RNA strands, which is responsible for translating RNA into protein inside cells.
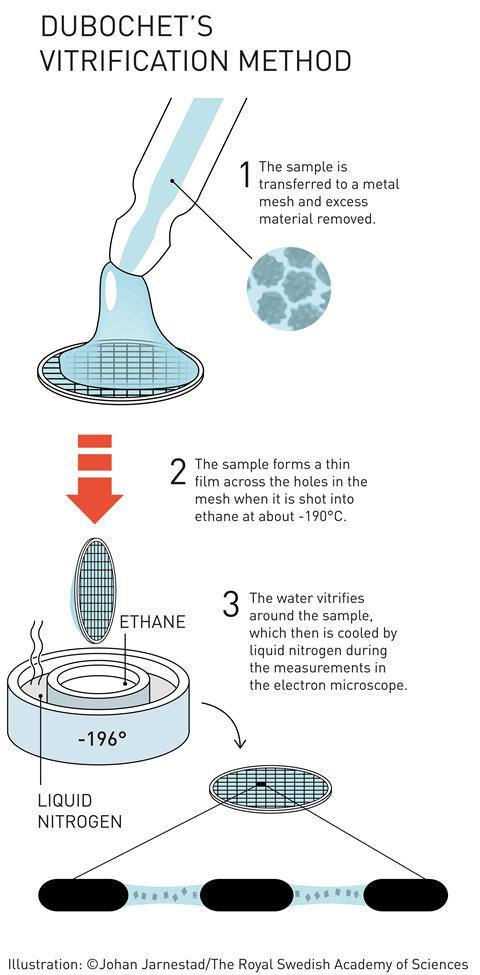
It was James Dubochet who put the ‘cryo’ into cryo-EM. He developed a method for freezing water-based TEM samples so rapidly that the water forms a disordered glass, rather than crystalline ice. This is important because ordered ice crystals would strongly diffract the microscope’s electron beam, obscuring any information about the molecules being studied. Catapulting the sample into a bath of liquid nitrogen-cooled ethane freezes the thin film of water on a TEM sample so quickly that the water molecules don’t have time to arrange into a crystalline lattice. In this ‘vitrified’ sample, the water is disordered but the 3D structure of the biomolecules in the sample is retained. Dubochet created the first images of various viruses using vitrified water samples.
What is cryo-EM used for?
Understanding how biomolecules function and interact is fundamental to biochemistry, and underpins efforts to develop new drugs and medical treatments, and understand and treat diseases.
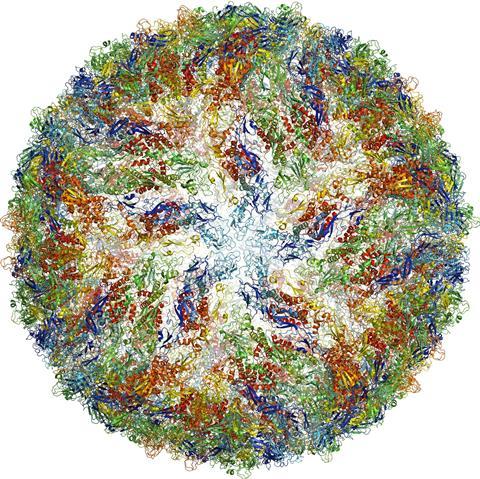
One recent example is the Zika virus. During the recent outbreak in Brazil, a group of researchers generated a high-resolution 3D image of the virus structure within a few months. This provided a starting point for searching for possible sites that could be targeted by drugs to prevent the spread of the virus.
More information on cryo-EM and the Nobel laureates can be found on the Nobel prize website














3 readers' comments