Protein that survived extreme conditions on early Earth could be a boon for biofuel industry
A 2.8-billion-year-old enzyme has been resurrected to help with industrial biofuel production. The cellulose-chewing protein, which evolved at the dawn of life when Earth was an inhospitable place, is at least twice as active as its modern counterparts under the harsh conditions of biomass breakdown.
Cellulose-containing materials – such as wood, cotton and even paper waste – are rich sources of chemicals, including biofuels like ethanol. To break down cellulose, the biochemical industry uses cellulase enzymes that convert the biopolymer into glucose. However, the conditions under which the raw materials are pretreated – high temperatures and strong acids – destroy cellulases.
Over the last two decades, says Raul Perez-Jimenez from Spain’s Nanoscience Cooperative Research Center, scientists have been trying to create more resistant cellulases – with limited success. His team has now taken a trip back in time to recreate an ancient enzyme that tolerates temperatures as high as 90°C and pH values between 4 and 10, while remaining at least twice as catalytically active as modern cellulases.
‘Instead of pushing evolution forward, we’re going backwards, look for those [proteins] that were once useful, but they were discarded as time passed,’ says Perez-Jimenez.
The idea of reconstructing ancient proteins isn’t new, it dates back to the 1960s. Since then, ancestral sequence reconstruction has grown into a tool that allows biologists to uncover evolutionary mechanisms. However, it has rarely been used to engineer enzymes and, according to Perez-Jimenez, never to improve several protein properties at once.
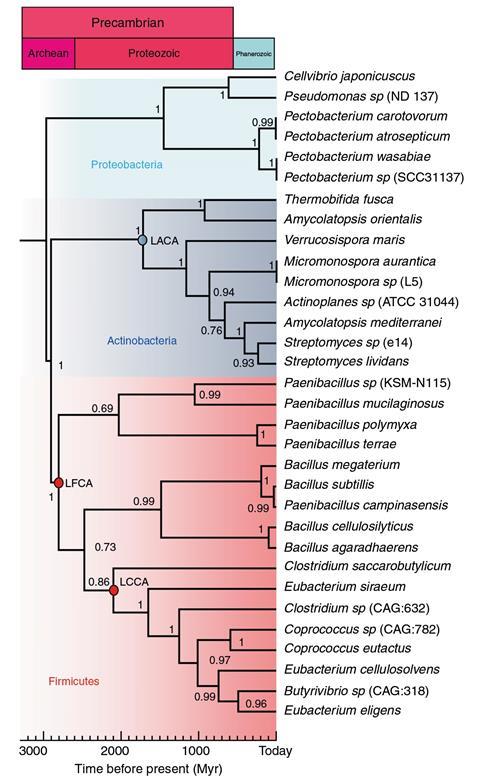
In ancestral sequence reconstruction, enzymes with the same function but coming from different species are used to construct a phylogenetic tree – a diagram showing evolutionary relationships. For each tree node, which represents the point where a common ancestor evolved into new species, researchers infer the most likely amino acid sequence according to the younger protein’s similarities and differences. Although this doesn’t perfectly reconstruct the extinct amino acid sequence, it can recreate the enzyme’s physical features, explains says Perez-Jimenez.
His team resurrected a cellulase from the Archean, 2.8 billion years ago. During this eon, the planet’s oceans were hot, acidic, salty and contained almost no oxygen. Archean organisms and their enzymes needed to be extremely hardy to survive. While there are some modern extremophiles that live in hot or acidic environments, very few can withstand both.
‘I thought it was [a] great [study],’ says evolutionary synthetic biologist Eric Gaucher from the Georgia Institute of Technology in the US. Other protein engineering methods – including directed evolution – create mostly random mutations, many of which aren’t functioning proteins, he explains. Since ancestral proteins are shaped by natural selection, they usually fold properly and retain their functionality.
While sequence reconstruction can’t create enzymes with completely new chemical functionality like directed evolution, Gaucher thinks that scientists will start to combine the two techniques. ‘Ancestral proteins can be a jumping off points for directed evolution experiments because they tend to be chemically more promiscuous than modern proteins. By going back in time, it’s like we’re going to the sandbox of evolution.’
‘We still see that many people [don’t] take this technique very seriously, more like a little bit of a science fiction,’ says Perez-Jimenez. ‘Our goal is to prove that this is a serious technology for enzyme design based on the principle of evolution.’
References
N Barruetabeña et al, Commun. Chem., 2019, 2, 76 (DOI: 10.1038/s42004-019-0176-6)














No comments yet