Scientists constructing enzyme library make serendipitous discovery
Researchers have made more active, heat tolerant versions of two potentially useful enzymes by reconstructing the forms thought to exist in the ancestor to all modern vertebrates 450 million years ago. They say this approach – demonstrated on a cytochrome P450 and a ketol-acid reductoisomerase (KARI) – could be useful for designing biocatalysts that are able to carry out desirable chemical transformations under industrial conditions.
Elizabeth Gillam at the University of Queensland in Australia explains cytochrome P450 enzymes, which are usually found in the liver and involved in metabolising various molecules, are of interest to chemists because they catalyse useful C–H functionalisations. But like other enzymes they tend to fall apart outside the body and don’t fare well at high temperatures. ‘What we’ve done is stabilise these biocatalysts,’ says Gillam. ‘We went back to the ancestral form of these enzymes, and it turns out that the ancestral form is very thermostable – it can take very high temperatures and it can last for a long time at ambient temperatures.’
Reconstructing ancestral enzymes isn’t an entirely new idea. Researchers have reasoned before that millions of years in the past, the Earth was much hotter, so enzymes that carried out similar reactions in the earliest life forms must have worked at higher temperatures. But while much work has focussed on the pre-Cambrian era (4500–540 million years ago), Gillam’s group reconstructed a more recent ancestral enzyme – that from the ancestor to all modern vertebrates.
‘The advantage of going only back to the earliest vertebrates is that we’ve got more confidence in the prediction of the ancestor’s sequence – the further back, the more uncertainty there is,’ explains Gillam.
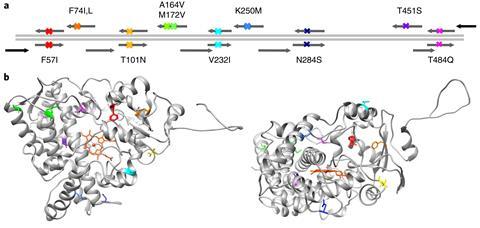
The team collected information on the genetic sequences of 138 different forms of the enzyme that exist in today’s vertebrates. ‘When you get all those sequences you align them – so you work out how different parts of the enzyme sequence align with other enzyme sequences – then from that you can derive an evolutionary tree,’ says Gillam. ‘With those two things – the alignment of sequences and the evolutionary tree – you can use bioinformatic algorithms to work out what the most likely ancestor was.’
The next step is to construct a gene for that enzyme, then introduce it into E. coli bacteria so that they will make the enzyme, which can be tested for properties like activity and stability. Gillam says the discovery that this particular ancestor had superior thermostability was serendipitous.
‘We didn’t actually approach this to derive a thermostable enzyme. What we were trying to do was just make a very large library of enzymes to try and map out what made an enzyme stable or able to be folded properly,’ she says. ‘It was a great surprise to us to find that it was very, very thermostable. We’d expect if you went back to pre-Cambrian conditions that an enzyme would be thermostable because the Earth went through times when the environment was very hot. But our ancestor was only from the earliest vertebrates … the temperatures around then were only about 10°C different than today.’
Hot topic
After realising the ancestral enzyme had these properties, the team used its sequence as a template for directed evolution, which allowed them to evolve enzymes that had even better thermal stability, capable of functioning at temperatures more than 30°C hotter and for around 100 times longer than modern cytochrome p450s.
They wanted to investigate whether the same approach would work with other enzymes, so they made the vertebrate ancestor enzyme of a KARI – enzymes which are currently used to make butanol-based biofuels. The ancestral form not only had superior thermostability, but was eight times more active than the E. coli form that is currently used as a model for biofuel production.
Paul Dalby, a biochemical engineer at University College London in the UK, praised the ‘impressive depth’ of the work, particularly the construction of libraries of variants on the enzymes’ possible ancestors. ‘They have shown that this could be a very good starting point for doing directed evolution,’ he adds.
References
Y Gumulya et al, Nat. Cat., 2018, DOI: 10.1038/s41929-018-0159-5















No comments yet